Non-targeted metabolomics unravels a media-dependent prodiginines production pathway in Streptomyces coelicolor A3(2)
- PMID: 30485320
- PMCID: PMC6261592
- DOI: 10.1371/journal.pone.0207541
Non-targeted metabolomics unravels a media-dependent prodiginines production pathway in Streptomyces coelicolor A3(2)
Abstract
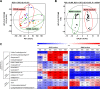
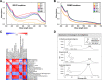
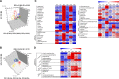
The genus Streptomyces is the best-known source of therapeutic secondary metabolites, especially antibiotics with pharmaceutical applications. Here, we performed a comparative study based on the time-resolved metabolic disparity in S. coelicolor A3(2) subjected to fermentative cultivation in two different types of media (R2YE and RSM3) in order to investigate secondary metabolite production pathways. The relative abundance of secondary metabolites, such as prodiginines, indoles, germicidins, and selected diketopiperazines, was increased in S. coelicolor A3(2) cultivated in R2YE medium compared to that in RSM3 medium, variably at the late-log and stationary phases of fermentative growth. Correlation analysis indicated that "antibiotic prodiginines" contributed maximally to the absorption maxima (A530) of culture supernatants, indicating their optimal production at 96 hours in R2YE medium. A higher abundance of L-proline (48-72 hours) followed by prodiginines (96 hours) was evident, substantiating the intertwined links between precursor and activated prodiginines pathway. Similarly, the higher abundance of indoles was concurrent with tryptophan levels in the shikimate pathway, whereas diketopiperazines were synchronously abundant along with the levels of phenylalanine, leucine, and proline. Additionally, acetyl-CoA induced the acetate pathway, resulting in the production of germicidins. Thus, our results demonstrate that S. coelicolor A3(2) produces specific secondary metabolites by enhancing the dedicated metabolic pathway responsible for their production. In conclusion, our results from this study provide insight into the metabolic pathways of S. coelicolor A3(2), and can be applied to further optimize the production of prodiginines.
Conflict of interest statement
Dynebio, Inc. provided the materials, for this study and is the employer of JHL, EJK, SJH, and YHL. There are no patents, products in development or marketed products to declare. This does not alter the authors' adherence to all the PLOS ONE policies on sharing data and materials, as detailed online in the guide for authors.
Figures


Similar articles
-
Multicopy proC in Streptomyces coelicolor A3(2) elicits a transient production of prodiginines, while proC deletion does not yield a proline auxotroph.J Mol Microbiol Biotechnol. 2010;19(3):152-8. doi: 10.1159/000321502. Epub 2010 Oct 6. J Mol Microbiol Biotechnol. 2010. PMID: 20924202
-
Production of branched-chain alkylprodiginines in S. coelicolor by replacement of the 3-ketoacyl ACP synthase III initiation enzyme, RedP.Chem Biol. 2005 Feb;12(2):191-200. doi: 10.1016/j.chembiol.2004.11.006. Chem Biol. 2005. PMID: 15734646
-
Optimized submerged batch fermentation strategy for systems scale studies of metabolic switching in Streptomyces coelicolor A3(2).BMC Syst Biol. 2012 Jun 7;6:59. doi: 10.1186/1752-0509-6-59. BMC Syst Biol. 2012. PMID: 22676814 Free PMC article.
-
New prodiginines from a ketosynthase swap.Chem Biol. 2005 Feb;12(2):145-6. doi: 10.1016/j.chembiol.2005.01.007. Chem Biol. 2005. PMID: 15734641 Review.
-
Molecular genetic analysis of proline and tryptophan biosynthesis in Streptomyces coelicolor A3(2): interaction between primary and secondary metabolism--a review.Gene. 1992 Jun 15;115(1-2):5-12. doi: 10.1016/0378-1119(92)90533-u. Gene. 1992. PMID: 1612450 Review.
Cited by
-
Improvement of Biosynthetic Ansamitocin P-3 Production Based on Oxygen-Vector Screening and Metabonomics Analysis.Evid Based Complement Alternat Med. 2022 Jun 2;2022:3564185. doi: 10.1155/2022/3564185. eCollection 2022. Evid Based Complement Alternat Med. 2022. PMID: 35692578 Free PMC article.
-
Multi-Omics Analysis of the Effect of cAMP on Actinorhodin Production in Streptomyces coelicolor.Front Bioeng Biotechnol. 2020 Nov 5;8:595552. doi: 10.3389/fbioe.2020.595552. eCollection 2020. Front Bioeng Biotechnol. 2020. PMID: 33251203 Free PMC article.
-
Natural Products Dereplication: Databases and Analytical Methods.Prog Chem Org Nat Prod. 2024;124:1-56. doi: 10.1007/978-3-031-59567-7_1. Prog Chem Org Nat Prod. 2024. PMID: 39101983 Review.
-
Continuous carbon source supply is essential for high rifamycin productivity of Amycolatopsis mediterranei in nitrate-stimulated fermentation revealed by a metabolomic study.Acta Biochim Biophys Sin (Shanghai). 2025 Jan 23;57(5):738-748. doi: 10.3724/abbs.2024245. Acta Biochim Biophys Sin (Shanghai). 2025. PMID: 39849912 Free PMC article.
-
Genomic and Untargeted Metabolomic Analysis of Secondary Metabolites in the Streptomyces griseoaurantiacus Strain MH191 Shows Media-Based Dependency for the Production of Bioactive Compounds with Potential Antifungal Activity.J Agric Food Chem. 2024 Nov 6;72(44):24432-24448. doi: 10.1021/acs.jafc.4c04989. Epub 2024 Oct 23. J Agric Food Chem. 2024. PMID: 39440812 Free PMC article.
References
-
- Berdy J. Bioactive microbial metabolites. J. Antibiot. 2005; 58(1): 1–26. 10.1038/ja.2005.1 - DOI - PubMed
-
- Hamaki T, Suzuki M, Fudou R., Jojima Y, Kajiura T, Tabuchi A, et al. Isolation of novel bacteria and actinomycetes using soil-extract agar medium. J Biosci Bioeng. 2005; 99(5):485–92. 10.1263/jbb.99.485 - DOI - PubMed
-
- Gomez-Escribano JP, Bibb MJ. Engineering Streptomyces coelicolor for heterologous expression of secondary metabolite gene clusters. Microb Biotechnol. 2011; 4(2):207–15. 10.1111/j.1751-7915.2010.00219.x - DOI - PMC - PubMed
-
- Venil CK, Zakaria ZA, Ahmad WA. Bacterial pigments and their applications. Process Biochem. 2013; 48:1065–1079. 10.1016/j.procbio.2013.06.006 - DOI
-
- Elibol M. Optimization of medium composition for actinorhodin production by Streptomyces coelicolor A3(2) with response surface methodology. Process Biochem. 2004; 39:1057–1062. 10.1016/S0032-9592(03)00232-2 - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

