The genomics of desmoplastic small round cell tumor reveals the deregulation of genes related to DNA damage response, epithelial-mesenchymal transition, and immune response
- PMID: 30486883
- PMCID: PMC6260689
- DOI: 10.1186/s40880-018-0339-3
The genomics of desmoplastic small round cell tumor reveals the deregulation of genes related to DNA damage response, epithelial-mesenchymal transition, and immune response
Abstract
Background: Desmoplastic small round cell tumor (DSRCT) is a rare, aggressive, and poorly investigated simple sarcoma with a low frequency of genetic deregulation other than an Ewing sarcoma RNA binding protein 1 (EWSR1)-Wilm's tumor suppressor (WT1) translocation. We used whole-exome sequencing to interrogate six consecutive pre-treated DSRCTs whose gene expression was previously investigated.
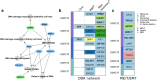
Methods: DNA libraries were prepared from formalin-fixed, paraffin-embedded archival tissue specimens following the Agilent SureSelectXT2 target enrichment protocol and sequenced on Illumina NextSeq 500. Raw sequence data were aligned to the reference genome with Burrows-Wheeler Aligner algorithm. Somatic mutations and copy number alterations (CNAs) were identified using MuTect2 and EXCAVATOR2, respectively. Biological functions associated with altered genes were investigated through Ingenuity Pathway Analysis (IPA) software.
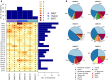
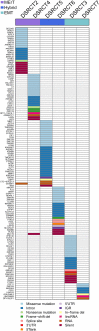
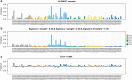
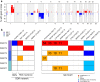
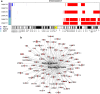
Results: A total of 137 unique somatic mutations were identified: 133 mutated genes were case-specific, and 2 were mutated in two cases but in different positions. Among the 135 mutated genes, 27% were related to two biological categories: DNA damage-response (DDR) network that was also identified through IPA and mesenchymal-epithelial reverse transition (MErT)/epithelial-mesenchymal transition (EMT) already demonstrated to be relevant in DSRCT. The mutated genes in the DDR network were involved in various steps of transcription and particularly affected pre-mRNA. Half of these genes encoded RNA-binding proteins or DNA/RNA-binding proteins, which were recently recognized as a new class of DDR players. CNAs in genes/gene families, involved in MErT/EMT and DDR, were recurrent across patients and mostly segregated in the MErT/EMT category. In addition, recurrent gains of regions in chromosome 1 involving many MErT/EMT gene families and loss of one arm or the entire chromosome 6 affecting relevant immune-regulatory genes were recorded.
Conclusions: The emerging picture is an extreme inter-tumor heterogeneity, characterized by the concurrent deregulation of the DDR and MErT/EMT dynamic and plastic programs that could favour genomic instability and explain the refractory DSRCT profile.
Keywords: Chromosome imbalance; Copy number alterations; DNA damage response; Desmoplastic small round cell tumor; Genomic stability; Immune response; Mesenchymal–epithelial reverse transition/epithelial–mesenchymal transition; Somatic mutations; Whole-exome sequencing.
Figures

Similar articles
-
Recurrent secondary genomic alterations in desmoplastic small round cell tumors.BMC Med Genet. 2020 May 11;21(1):101. doi: 10.1186/s12881-020-01034-w. BMC Med Genet. 2020. PMID: 32393201 Free PMC article.
-
New transcriptional-based insights into the pathogenesis of desmoplastic small round cell tumors (DSRCTs).Oncotarget. 2017 May 16;8(20):32492-32504. doi: 10.18632/oncotarget.16477. Oncotarget. 2017. PMID: 28415643 Free PMC article.
-
A genomic case study of desmoplastic small round cell tumor: comprehensive analysis reveals insights into potential therapeutic targets and development of a monitoring tool for a rare and aggressive disease.Hum Genomics. 2016 Nov 18;10(1):36. doi: 10.1186/s40246-016-0092-0. Hum Genomics. 2016. PMID: 27863505 Free PMC article.
-
Mini-Review on Targeted Treatment of Desmoplastic Small Round Cell Tumor.Front Oncol. 2020 Apr 21;10:518. doi: 10.3389/fonc.2020.00518. eCollection 2020. Front Oncol. 2020. PMID: 32373525 Free PMC article. Review.
-
Desmoplastic small round cell tumor of the parotid gland-report of a rare case and a review of the literature.Diagn Pathol. 2019 May 18;14(1):43. doi: 10.1186/s13000-019-0825-1. Diagn Pathol. 2019. PMID: 31103034 Free PMC article. Review.
Cited by
-
Genomic Breakpoint Characterization and Transcriptome Analysis of Metastatic, Recurrent Desmoplastic Small Round Cell Tumor.Sarcoma. 2023 Jul 6;2023:6686702. doi: 10.1155/2023/6686702. eCollection 2023. Sarcoma. 2023. PMID: 37457440 Free PMC article.
-
Olaparib and temozolomide in desmoplastic small round cell tumors: a promising combination in vitro and in vivo.J Cancer Res Clin Oncol. 2020 Jul;146(7):1659-1670. doi: 10.1007/s00432-020-03211-z. Epub 2020 Apr 11. J Cancer Res Clin Oncol. 2020. PMID: 32279088 Free PMC article.
-
Comprehensive Molecular Profiling of Desmoplastic Small Round Cell Tumor.Mol Cancer Res. 2021 Jul;19(7):1146-1155. doi: 10.1158/1541-7786.MCR-20-0722. Epub 2021 Mar 22. Mol Cancer Res. 2021. PMID: 33753552 Free PMC article.
-
Recurrent secondary genomic alterations in desmoplastic small round cell tumors.BMC Med Genet. 2020 May 11;21(1):101. doi: 10.1186/s12881-020-01034-w. BMC Med Genet. 2020. PMID: 32393201 Free PMC article.
-
Pediatric sarcomas display a variable EpCAM expression in a histology-dependent manner.Transl Oncol. 2020 Nov;13(11):100846. doi: 10.1016/j.tranon.2020.100846. Epub 2020 Aug 14. Transl Oncol. 2020. PMID: 32805674 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials

