Distributive enzyme binding controlled by local RNA context results in 3' to 5' directional processing of dicistronic tRNA precursors by Escherichia coli ribonuclease P
- PMID: 30496557
- PMCID: PMC6379654
- DOI: 10.1093/nar/gky1162
Distributive enzyme binding controlled by local RNA context results in 3' to 5' directional processing of dicistronic tRNA precursors by Escherichia coli ribonuclease P
Abstract
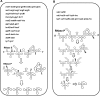
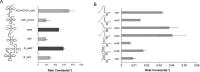
RNA processing by ribonucleases and RNA modifying enzymes often involves sequential reactions of the same enzyme on a single precursor transcript. In Escherichia coli, processing of polycistronic tRNA precursors involves separation into individual pre-tRNAs by one of several ribonucleases followed by 5' end maturation by ribonuclease P. A notable exception are valine and lysine tRNAs encoded by three polycistronic precursors that follow a recently discovered pathway involving initial 3' to 5' directional processing by RNase P. Here, we show that the dicistronic precursor containing tRNAvalV and tRNAvalW undergoes accurate and efficient 3' to 5' directional processing by RNase P in vitro. Kinetic analyses reveal a distributive mechanism involving dissociation of the enzyme between the two cleavage steps. Directional processing is maintained despite swapping or duplicating the two tRNAs consistent with inhibition of processing by 3' trailer sequences. Structure-function studies identify a stem-loop in 5' leader of tRNAvalV that inhibits RNase P cleavage and further enforces directional processing. The results demonstrate that directional processing is an intrinsic property of RNase P and show how RNA sequence and structure context can modulate reaction rates in order to direct precursors along specific pathways.
Published by Oxford University Press on behalf of Nucleic Acids Research 2018.
Figures
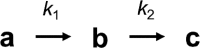







References
-
- Blattner F.R., Plunkett G. 3rd, Bloch C.A., Perna N.T., Burland V., Riley M., Collado-Vides J., Glasner J.D., Rode C.K., Mayhew G.F. et al. The complete genome sequence of Escherichia coli K-12. Science. 1997; 277:1453–1462. - PubMed
-
- Li Z., Deutscher M.P.. Maturation pathways for E. coli tRNA precursors: a random multienzyme process in vivo. Cell. 1996; 86:503–512. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

