Construction of a highly saturated linkage map in Japanese plum (Prunus salicina L.) using GBS for SNP marker calling
- PMID: 30507961
- PMCID: PMC6277071
- DOI: 10.1371/journal.pone.0208032
Construction of a highly saturated linkage map in Japanese plum (Prunus salicina L.) using GBS for SNP marker calling
Abstract
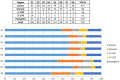
This study reports the construction of high density linkage maps of Japanese plum (Prunus salicina Lindl.) using single nucleotide polymorphism markers (SNPs), obtained with a GBS strategy. The mapping population (An x Au) was obtained by crossing cv. "Angeleno" (An) as maternal line and cv. "Aurora" (Au) as the pollen donor. A total of 49,826 SNPs were identified using the peach genome V2.1 as a reference. Then a stringent filtering was carried out, which revealed 1,441 high quality SNPs in 137 An x Au offspring, which were mapped in eight linkage groups. Finally, the consensus map was built using 732 SNPs which spanned 617 cM with an average of 0.96 cM between adjacent markers. The majority of the SNPs were distributed in the intragenic region in all the linkage groups. Considering all linkage groups together, 85.6% of the SNPs were located in intragenic regions and only 14.4% were located in intergenic regions. The genetic linkage analysis was able to co-localize two to three SNPs over 37 putative orthologous genes in eight linkage groups in the Japanese plum map. These results indicate a high level of synteny and collinearity between Japanese plum and peach genomes.
Conflict of interest statement
Author Rolando García-González is employed by Sociedad BioTECNOS Ltda. This affiliation commenced towards the end of this study. There are no patents, products in development or marketed products to declare. This does not alter our adherence to all the PLOS ONE policies on sharing data and materials.
Figures


Similar articles
-
Construction of High Density Sweet Cherry (Prunus avium L.) Linkage Maps Using Microsatellite Markers and SNPs Detected by Genotyping-by-Sequencing (GBS).PLoS One. 2015 May 26;10(5):e0127750. doi: 10.1371/journal.pone.0127750. eCollection 2015. PLoS One. 2015. PMID: 26011256 Free PMC article.
-
High-density multi-population consensus genetic linkage map for peach.PLoS One. 2018 Nov 21;13(11):e0207724. doi: 10.1371/journal.pone.0207724. eCollection 2018. PLoS One. 2018. PMID: 30462743 Free PMC article.
-
Genotyping by Sequencing for SNP-Based Linkage Analysis and Identification of QTLs Linked to Fruit Quality Traits in Japanese Plum (Prunus salicina Lindl.).Front Plant Sci. 2017 Apr 11;8:476. doi: 10.3389/fpls.2017.00476. eCollection 2017. Front Plant Sci. 2017. PMID: 28443103 Free PMC article.
-
A high-density SNP-based linkage map using genotyping-by-sequencing and its utilization for improved genome assembly of chickpea (Cicer arietinum L.).Funct Integr Genomics. 2020 Nov;20(6):763-773. doi: 10.1007/s10142-020-00751-y. Epub 2020 Aug 27. Funct Integr Genomics. 2020. PMID: 32856221
-
Current achievements and future directions in genetic engineering of European plum (Prunus domestica L.).Transgenic Res. 2018 Jun;27(3):225-240. doi: 10.1007/s11248-018-0072-3. Epub 2018 Apr 12. Transgenic Res. 2018. PMID: 29651659 Free PMC article. Review.
Cited by
-
QTLs Identification for Iron Chlorosis in a Segregating Peach-Almond Progeny Through Double-Digest Sequence-Based Genotyping (SBG).Front Plant Sci. 2022 May 31;13:872208. doi: 10.3389/fpls.2022.872208. eCollection 2022. Front Plant Sci. 2022. PMID: 35712560 Free PMC article.
-
Chromosome-level draft genome of a diploid plum (Prunus salicina).Gigascience. 2020 Dec 10;9(12):giaa130. doi: 10.1093/gigascience/giaa130. Gigascience. 2020. PMID: 33300949 Free PMC article.
-
Classification of Takifugu rubripes, T. chinensis and T. pseudommus by genotyping-by-sequencing.PLoS One. 2020 Aug 27;15(8):e0236483. doi: 10.1371/journal.pone.0236483. eCollection 2020. PLoS One. 2020. PMID: 32853203 Free PMC article.
-
Descriptive Genomic Analysis and Sequence Genotyping of the Two Papaya Species (Vasconcellea pubescens and Vasconcellea chilensis) Using GBS Tools.Plants (Basel). 2022 Aug 18;11(16):2151. doi: 10.3390/plants11162151. Plants (Basel). 2022. PMID: 36015454 Free PMC article.
-
Advances in genomics for diversity studies and trait improvement in temperate fruit and nut crops under changing climatic scenarios.Front Plant Sci. 2023 Jan 19;13:1048217. doi: 10.3389/fpls.2022.1048217. eCollection 2022. Front Plant Sci. 2023. PMID: 36743560 Free PMC article. Review.
References
-
- Carrasco B, Meisel L, Gebauer M, Gonzalez RG, Silva H. Breeding in peach, cherry and plum: from a tissue culture, genetic, transcriptomic and genomic perspective. Biological Research. 2013;46(3):219–30. 10.4067/S0716-97602013000300001 - DOI - PubMed
-
- Okie WR, Weinberger JH. Plums. In: Janick J, Moore JN, editors. Fruit breeding, Tree and Tropical Fruits, 2nd ed New York: John Wiley & Sons; 1995. p. 559–607.
-
- Hancock JF. Temperate fruit crop breeding: germplasm to genomics. Hancock JF, editor. Netherland: Springer Science & Business Media; 2008. 1–455 p.
-
- El-Sharkawy I, Kim WS, El-Kereamy A, Jayasankar S, Svircev AM, Brown DCW. Isolation and characterization of four ethylene signal transduction elements in plums (Prunus salicina L.). Journal of experimental botany. 2007;58(13):3631–43. 10.1093/jxb/erm213 - DOI - PubMed
-
- El-Sharkawy I, Sherif S, Mila I, Bouzayen M, Jayasankar S. Molecular characterization of seven genes encoding ethylene-responsive transcriptional factors during plum fruit development and ripening. Journal of experimental botany. 2009;60(3):907–22. 10.1093/jxb/ern354 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

