ClinPhen extracts and prioritizes patient phenotypes directly from medical records to expedite genetic disease diagnosis
- PMID: 30514889
- PMCID: PMC6551315
- DOI: 10.1038/s41436-018-0381-1
ClinPhen extracts and prioritizes patient phenotypes directly from medical records to expedite genetic disease diagnosis
Abstract
Purpose: Diagnosing monogenic diseases facilitates optimal care, but can involve the manual evaluation of hundreds of genetic variants per case. Computational tools like Phrank expedite this process by ranking all candidate genes by their ability to explain the patient's phenotypes. To use these tools, busy clinicians must manually encode patient phenotypes from lengthy clinical notes. With 100 million human genomes estimated to be sequenced by 2025, a fast alternative to manual phenotype extraction from clinical notes will become necessary.
Methods: We introduce ClinPhen, a fast, high-accuracy tool that automatically converts clinical notes into a prioritized list of patient phenotypes using Human Phenotype Ontology (HPO) terms.
Results: ClinPhen shows superior accuracy and 20× speedup over existing phenotype extractors, and its novel phenotype prioritization scheme improves the performance of gene-ranking tools.
Conclusion: While a dedicated clinician can process 200 patient records in a 40-hour workweek, ClinPhen does the same in 10 minutes. Compared with manual phenotype extraction, ClinPhen saves an additional 3-5 hours per Mendelian disease diagnosis. Providers can now add ClinPhen's output to each summary note attached to a filled testing laboratory request form. ClinPhen makes a substantial contribution to improvements in efficiency critically needed to meet the surging demand for clinical diagnostic sequencing.
Keywords: Mendelian disease diagnosis; medical genetics; natural language processing; prioritized disease phenotypes.
Conflict of interest statement
DISCLOSURE
The authors declare no conflicts of interest.
Figures
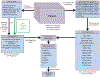



References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

