Plasma donor-derived cell-free DNA kinetics after kidney transplantation using a single tube multiplex PCR assay
- PMID: 30521549
- PMCID: PMC6283554
- DOI: 10.1371/journal.pone.0208207
Plasma donor-derived cell-free DNA kinetics after kidney transplantation using a single tube multiplex PCR assay
Abstract
Background: After transplantation, cell-free DNA derived from the donor organ (ddcfDNA) can be detected in the recipient's circulation. We aimed to quantify ddcfDNA levels in plasma of kidney transplant recipients thereby investigating the kinetics of this biomarker after transplantation and determining biological variables that influence ddcfDNA kinetics in stable and non-stable patients.
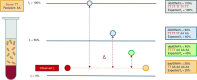
Materials and methods: From 107 kidney transplant recipients, plasma samples were collected longitudinally after transplantation (day 1-3 months) within a multicenter set-up. Cell-free DNA from the donor was quantified in plasma as a fraction of the total cell-free DNA by next generation sequencing using a targeted, multiplex PCR-based method for the analysis of single nucleotide polymorphisms. A subgroup of stable renal transplant recipients was identified to determine a ddcfDNA threshold value.
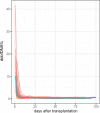
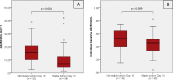
Results: In stable transplant recipients, plasma ddcfDNA% decreased to a mean (SD) ddcfDNA% of 0.46% (± 0.21%) which was reached 9.85 (± 5.6) days after transplantation. A ddcfDNA threshold value of 0.88% (mean + 2SD) was determined in kidney transplant recipients. Recipients that did not reach this threshold ddcfDNA value within 10 days after transplantation showed a higher ddcfDNA% on the first day after transplantation and demonstrated a higher individual baseline ddcfDNA%.
Conclusion: In conclusion, plasma ddcfDNA fractions decreased exponentially within 10 days after transplantation to a ddcfDNA threshold value of 0.88% or less. To investigate the role of ddcfDNA for rejection monitoring of the graft, future research is needed to determine causes of ddcfDNA% increases above this threshold value.
Conflict of interest statement
We have read the journal's policy and the authors of this manuscript have added the following competing interests: Jurgen Del Favero and Joachim De Schrijver are affiliated to Multiplicom N.V. This affiliation does not alter our adherence to PLOS ONE policies on sharing data and materials.
Figures


References
-
- Gielis EM, Ledeganck KJ, De Winter BY, Del Favero J, Bosmans JL, Claas FH, et al. Cell-Free DNA: An Upcoming Biomarker in Transplantation. American journal of transplantation: official journal of the American Society of Transplantation and the American Society of Transplant Surgeons. 2015;15(10):2541–51. - PubMed
-
- De Vlaminck I, Valantine HA, Snyder TM, Strehl C, Cohen G, Luikart H, et al. Circulating cell-free DNA enables noninvasive diagnosis of heart transplant rejection. Science translational medicine. 2014;6(241):241ra77 10.1126/scitranslmed.3007803 - DOI - PMC - PubMed
-
- Schutz E, Fischer A, Beck J, Harden M, Koch M, Wuensch T, et al. Graft-derived cell-free DNA, a noninvasive early rejection and graft damage marker in liver transplantation: A prospective, observational, multicenter cohort study. PLoS medicine. 2017;14(4):e1002286 10.1371/journal.pmed.1002286 - DOI - PMC - PubMed
-
- De Vlaminck I, Martin L, Kertesz M, Patel K, Kowarsky M, Strehl C, et al. Noninvasive monitoring of infection and rejection after lung transplantation. Proceedings of the National Academy of Sciences of the United States of America. 2015;112(43):13336–41. 10.1073/pnas.1517494112 - DOI - PMC - PubMed
-
- Lo YM, Tein MS, Pang CC, Yeung CK, Tong KL, Hjelm NM. Presence of donor-specific DNA in plasma of kidney and liver-transplant recipients. Lancet. 1998;351(9112):1329–30. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

