lncITPF Promotes Pulmonary Fibrosis by Targeting hnRNP-L Depending on Its Host Gene ITGBL1
- PMID: 30528088
- PMCID: PMC6369732
- DOI: 10.1016/j.ymthe.2018.08.026
lncITPF Promotes Pulmonary Fibrosis by Targeting hnRNP-L Depending on Its Host Gene ITGBL1
Abstract
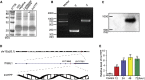
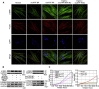
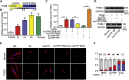
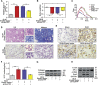
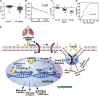
The role of long non-coding RNA (lncRNA) in idiopathic pulmonary fibrosis (IPF) is poorly understood. We found a novel lncRNA-ITPF that was upregulated in IPF. Bioinformatics and in vitro translation verified that lncITPF is an actual lncRNA, and its conservation is in evolution. Northern blot and rapid amplification of complementary DNA ends were used to analyze the full-length sequence of lncITPF. RNA fluorescence in situ hybridization and nucleocytoplasmic separation demonstrated that lncITPF was mainly located in the nucleus. RNA sequencing, chromatin immunoprecipitation (ChIP)-qPCR, CRISPR-Cas9 technology, and promoter activity analysis showed that the fibrotic function of lncITPF depends on its host gene integrin β-like 1 (ITGBL1), but they did not share the same promoter and were not co-transcribed. Luciferase activity, pathway inhibitors, and ChIP-qPCR showed that smad2/3 binds to the lncITPF promoter, and TGF-β1-smad2/3 was the upstream inducer of the fibrotic pathway. Furthermore, RNA-protein pull-down, liquid chromatography-mass spectrometry (LC-MS), and protein-RNA immunoprecipitation showed that lncITPF regulated H3 and H4 histone acetylation in the ITGBL1 promoter by targeting heterogeneous nuclear ribonucleoprotein L. Finally, sh-lncITPF was used to evaluate the therapeutic effect of lncITPF. Clinical analysis showed that lncITPF is associated with the clinicopathological features of IPF patients. Our findings provide a therapeutic target or diagnostic biomarker for IPF.
Keywords: CRISPR-Cas9 technology; IPF; ITGBL1; RNA-protein pull-down; RNA-sequencing; TGF-β1-smad2/3; lncITPF; lncRNA.
Copyright © 2018. Published by Elsevier Inc.
Figures



Similar articles
-
Astaxanthin attenuates pulmonary fibrosis through lncITPF and mitochondria-mediated signal pathways.J Cell Mol Med. 2020 Sep;24(17):10245-10250. doi: 10.1111/jcmm.15477. Epub 2020 Aug 19. J Cell Mol Med. 2020. PMID: 32813323 Free PMC article.
-
MOBT Alleviates Pulmonary Fibrosis through an lncITPF-hnRNP-l-Complex-Mediated Signaling Pathway.Molecules. 2022 Aug 22;27(16):5336. doi: 10.3390/molecules27165336. Molecules. 2022. PMID: 36014574 Free PMC article.
-
A novel lnc-PCF promotes the proliferation of TGF-β1-activated epithelial cells by targeting miR-344a-5p to regulate map3k11 in pulmonary fibrosis.Cell Death Dis. 2017 Oct 26;8(10):e3137. doi: 10.1038/cddis.2017.500. Cell Death Dis. 2017. PMID: 29072702 Free PMC article.
-
Knockdown of Long Noncoding RNA H19 Represses the Progress of Pulmonary Fibrosis through the Transforming Growth Factor β/Smad3 Pathway by Regulating MicroRNA 140.Mol Cell Biol. 2019 May 28;39(12):e00143-19. doi: 10.1128/MCB.00143-19. Print 2019 Jun 15. Mol Cell Biol. 2019. PMID: 30988156 Free PMC article.
-
The Long Noncoding RNA DNM3OS Is a Reservoir of FibromiRs with Major Functions in Lung Fibroblast Response to TGF-β and Pulmonary Fibrosis.Am J Respir Crit Care Med. 2019 Jul 15;200(2):184-198. doi: 10.1164/rccm.201807-1237OC. Am J Respir Crit Care Med. 2019. PMID: 30964696
Cited by
-
LncRNA Hoxaas3 promotes lung fibroblast activation and fibrosis by targeting miR-450b-5p to regulate Runx1.Cell Death Dis. 2020 Aug 26;11(8):706. doi: 10.1038/s41419-020-02889-w. Cell Death Dis. 2020. PMID: 32848140 Free PMC article.
-
Danshensu methyl ester enhances autophagy to attenuate pulmonary fibrosis by targeting lncIAPF-HuR complex.Front Pharmacol. 2022 Oct 31;13:1013098. doi: 10.3389/fphar.2022.1013098. eCollection 2022. Front Pharmacol. 2022. PMID: 36386240 Free PMC article.
-
Long non-coding RNAs: Promising new targets in pulmonary fibrosis.J Gene Med. 2021 Mar;23(3):e3318. doi: 10.1002/jgm.3318. Epub 2021 Feb 11. J Gene Med. 2021. PMID: 33533071 Free PMC article. Review.
-
Astaxanthin attenuates pulmonary fibrosis through lncITPF and mitochondria-mediated signal pathways.J Cell Mol Med. 2020 Sep;24(17):10245-10250. doi: 10.1111/jcmm.15477. Epub 2020 Aug 19. J Cell Mol Med. 2020. PMID: 32813323 Free PMC article.
-
Long Non-Coding RNAs in Cardiac and Pulmonary Fibroblasts and Fibrosis.Noncoding RNA. 2022 Jul 15;8(4):53. doi: 10.3390/ncrna8040053. Noncoding RNA. 2022. PMID: 35893236 Free PMC article.
References
-
- King T.E., Jr., Albera C., Bradford W.Z., Costabel U., du Bois R.M., Leff J.A., Nathan S.D., Sahn S.A., Valeyre D., Noble P.W. All-cause mortality rate in patients with idiopathic pulmonary fibrosis. Implications for the design and execution of clinical trials. Am. J. Respir. Crit. Care Med. 2014;189:825–831. - PubMed
-
- King T.E., Jr., Pardo A., Selman M. Idiopathic pulmonary fibrosis. Lancet. 2011;378:1949–1961. - PubMed
-
- Liu Y.M., Nepali K., Liou J.P. Idiopathic Pulmonary Fibrosis: Current Status, Recent Progress, and Emerging Targets. J. Med. Chem. 2017;60:527–553. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

