Detection and Analysis of the Hedgehog Signaling Pathway-Related Long Non-Coding RNA (lncRNA) Expression Profiles in Keloid
- PMID: 30543583
- PMCID: PMC6301256
- DOI: 10.12659/MSM.911159
Detection and Analysis of the Hedgehog Signaling Pathway-Related Long Non-Coding RNA (lncRNA) Expression Profiles in Keloid
Abstract
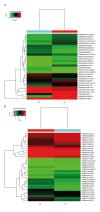
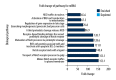
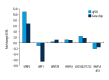
BACKGROUND Hedgehog (Hh) signaling pathway-related genes have important roles in several physiological and disease processes that involve cell proliferation. Long non-coding region RNAs (lncRNAs) have a regulatory role on gene expression. Keloid is characterized by excessive proliferation of scar tissue following trauma. The aims of this study were to evaluate the Hh signaling pathway in keloid skin tissues and its downstream gene expression and lncRNAs, compared with normal skin. MATERIAL AND METHODS Four pairs of keloids and adjacent normal skin epidermis underwent total RNA extraction. Gene chip high-throughput real-time quantitative polymerase chain reaction (qPCR) was used to examine the differential expression profiles of the Hh signaling pathway-related lncRNAs and mRNAs in the human keloid and normal skin. The differentially expressed mRNAs were analyzed by Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) to identify their biological roles. RESULTS In keloid tissue, differential expression of 33 mRNAs and 30 lncRNAs relating to the Hh pathway, were verified by gene chip qPCR. The results of GO and KEGG analysis showed that the upregulated mRNAs were involved in cell proliferation, cell growth, and tissue repair, and down-regulated mRNAs were involved in apoptosis. The lncRNA, AC073257.2, affected cell keloid growth and proliferation by its upstream target the GLI2 gene at the transcriptional level. The lncRNA, HNF1A-AS1, affected cell keloid growth and proliferation by its neighboring target gene, HNF1A. CONCLUSIONS Differential expression occurred in Hh signaling pathway-related lncRNAs and mRNAs, which may provide further insight into the development of keloid.
Figures

References
-
- Profyris C, Tziotzios C, Do Vale I. Cutaneous scarring: Pathophysiology, molecular mechanisms, and scar reduction therapeutics Part I. The molecular basis of scar formation. J Am Acad Dermatol. 2012;66:1–12. - PubMed
-
- Butler PD, Longaker MT, Yang GP. Current progress in keloid research and treatment. J Am Coll Surg. 2008;206:731–41. - PubMed
-
- Zhao L, Kong H, Sun H, et al. LncRNA-PVT1 promotes pancreatic cancer cells proliferation and migration through acting as a molecular sponge to regulate miR-448. J Cell Physiol. 2017;10:1002. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

