Comprehensive identification of protein disulfide bonds with pepsin/trypsin digestion, Orbitrap HCD and Spectrum Identification Machine
- PMID: 30557666
- PMCID: PMC6414265
- DOI: 10.1016/j.jprot.2018.12.010
Comprehensive identification of protein disulfide bonds with pepsin/trypsin digestion, Orbitrap HCD and Spectrum Identification Machine
Abstract
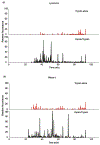
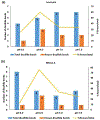
Disulfide bonds (SS) are post-translational modifications important for the proper folding and stabilization of many cellular proteins with therapeutic uses, including antibodies and other biologics. With budding advances of biologics and biosimilars, there is a mounting need for a robust method for accurate identification of SS. Even though several mass spectrometry methods have emerged for this task, their practical use rests on the broad effectiveness of both sample preparation methods and bioinformatics tools. Here we present a new protocol tailored toward mapping SS; it uses readily available reagents, instruments, and software. For sample preparation, a 4-h pepsin digestion at pH 1.3 followed by an overnight trypsin digestion at pH 6.5 can maximize the release of SS-containing peptides from non-reduced proteins, while minimizing SS scrambling. For LC/MS/MS analysis, SS-containing peptides can be efficiently fragmented with HCD in a Q Exactive Orbitrap mass spectrometer, preserving SS for subsequent identification. Our bioinformatics protocol describes how we tailored our freely downloadable and easy-to-use software, Spectrum Identification Machine for Cross-Linked Peptides (SIM-XL), to minimize false identification and facilitate manual validation of SS-peptide mass spectra. To substantiate this optimized method, we've comprehensively identified 14 out of 17 known SS in BSA. SIGNIFICANCE: Comprehensive and accurate identification of SS in proteins is critical for elucidating protein structures and functions. Yet, it is far from routine to accomplish this task in many analytical or core laboratories. Numerous published methods require complex sample preparation methods, specialized mass spectrometers and cumbersome or proprietary software tools, thus cannot be easily implemented in unspecialized laboratories. Here, we describe a robust and rapid SS mapping approach that utilizes readily available reagents, instruments, and software; it can be easily implemented in any analytical core laboratories, and tested for its impact on the research community.
Keywords: HCD; Pepsin; Protein disulfide bond; SIM-XL; Tandem mass spectrometry; Trypsin.
Copyright © 2018 Elsevier B.V. All rights reserved.
Conflict of interest statement
Conflict of interest
The authors declare that there is no conflict of interest.
Figures



Similar articles
-
Facilitating protein disulfide mapping by a combination of pepsin digestion, electron transfer higher energy dissociation (EThcD), and a dedicated search algorithm SlinkS.Mol Cell Proteomics. 2014 Oct;13(10):2776-86. doi: 10.1074/mcp.O114.039057. Epub 2014 Jun 30. Mol Cell Proteomics. 2014. PMID: 24980484 Free PMC article.
-
Development of a sample preparation method for monitoring correct disulfide linkages of monoclonal antibodies by liquid chromatography-mass spectrometry.Anal Biochem. 2016 Feb 15;495:21-8. doi: 10.1016/j.ab.2015.11.010. Epub 2015 Nov 30. Anal Biochem. 2016. PMID: 26656925
-
In-Depth Characterization of Protein Disulfide Bonds by Online Liquid Chromatography-Electrochemistry-Mass Spectrometry.J Am Soc Mass Spectrom. 2016 Jan;27(1):50-8. doi: 10.1007/s13361-015-1258-z. Epub 2015 Sep 14. J Am Soc Mass Spectrom. 2016. PMID: 26369777 Free PMC article.
-
Protein disulfide bond determination by mass spectrometry.Mass Spectrom Rev. 2002 May-Jun;21(3):183-216. doi: 10.1002/mas.10025. Mass Spectrom Rev. 2002. PMID: 12476442 Review.
-
Recent mass spectrometry-based techniques and considerations for disulfide bond characterization in proteins.Anal Bioanal Chem. 2018 Apr;410(10):2467-2484. doi: 10.1007/s00216-017-0772-1. Epub 2017 Dec 18. Anal Bioanal Chem. 2018. PMID: 29256076 Free PMC article. Review.
Cited by
-
Characterization of the Interactions between Minocycline Hydrochloride and Trypsin with Spectroscopic and Molecular Docking Technology.Molecules. 2023 Mar 15;28(6):2656. doi: 10.3390/molecules28062656. Molecules. 2023. PMID: 36985629 Free PMC article.
-
Selective cysteines oxidation in soluble guanylyl cyclase catalytic domain is involved in NO activation.Free Radic Biol Med. 2021 Jan;162:450-460. doi: 10.1016/j.freeradbiomed.2020.11.001. Epub 2020 Nov 6. Free Radic Biol Med. 2021. PMID: 33161042 Free PMC article.
-
Comprehensive Analysis of Tryptic Peptides Arising from Disulfide Linkages in NISTmAb and Their Use for Developing a Mass Spectral Library.J Proteome Res. 2021 Mar 5;20(3):1612-1629. doi: 10.1021/acs.jproteome.0c00823. Epub 2021 Feb 8. J Proteome Res. 2021. PMID: 33555887 Free PMC article.
-
Structural plasticity of the coiled-coil interactions in human SFPQ.Nucleic Acids Res. 2025 Jan 11;53(2):gkae1198. doi: 10.1093/nar/gkae1198. Nucleic Acids Res. 2025. PMID: 39698821 Free PMC article.
References
-
- Hogg PJ, Disulfide bonds as switches for protein function, Trends Biochem Sci 28(4) (2003) 210–4. - PubMed
-
- Thornton JM, Disulphide bridges in globular proteins, J Mol Biol 151(2) (1981) 261–87. - PubMed
-
- Nakamura T, Lipton SA, Cell death: protein misfolding and neurodegenerative diseases, Apoptosis 14(4) (2009) 455–68. - PubMed
-
- Dranoff G, Targets of protective tumor immunity, Ann N Y Acad Sci 1174 (2009) 74–80. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

