Regulation of Structure-Specific Endonucleases in Replication Stress
- PMID: 30558228
- PMCID: PMC6316474
- DOI: 10.3390/genes9120634
Regulation of Structure-Specific Endonucleases in Replication Stress
Abstract
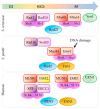
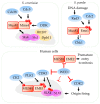
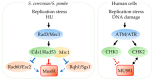
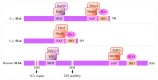
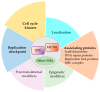
Replication stress results in various forms of aberrant replication intermediates that need to be resolved for faithful chromosome segregation. Structure-specific endonucleases (SSEs) recognize DNA secondary structures rather than primary sequences and play key roles during DNA repair and replication stress. Holliday junction resolvase MUS81 (methyl methane sulfonate (MMS), and UV-sensitive protein 81) and XPF (xeroderma pigmentosum group F-complementing protein) are a subset of SSEs that resolve aberrant replication structures. To ensure genome stability and prevent unnecessary DNA breakage, these SSEs are tightly regulated by the cell cycle and replication checkpoints. We discuss the regulatory network that control activities of MUS81 and XPF and briefly mention other SSEs involved in the resolution of replication intermediates.
Keywords: Mus81; XPF; replication stress; structure-specific endonuclease.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Control of structure-specific endonucleases to maintain genome stability.Nat Rev Mol Cell Biol. 2017 May;18(5):315-330. doi: 10.1038/nrm.2016.177. Epub 2017 Mar 22. Nat Rev Mol Cell Biol. 2017. PMID: 28327556 Review.
-
Structure-Specific Endonucleases and the Resolution of Chromosome Underreplication.Genes (Basel). 2019 Mar 19;10(3):232. doi: 10.3390/genes10030232. Genes (Basel). 2019. PMID: 30893921 Free PMC article. Review.
-
The Structure-Specific Endonucleases MUS81 and SEND1 Are Essential for Telomere Stability in Arabidopsis.Plant Cell. 2016 Jan;28(1):74-86. doi: 10.1105/tpc.15.00898. Epub 2015 Dec 24. Plant Cell. 2016. PMID: 26704385 Free PMC article.
-
Subnuclear Relocalization of Structure-Specific Endonucleases in Response to DNA Damage.Cell Rep. 2017 Aug 15;20(7):1553-1562. doi: 10.1016/j.celrep.2017.07.059. Cell Rep. 2017. PMID: 28813668
-
Analysis of Structure-Selective Endonuclease Activities From Yeast and Human Extracts.Methods Enzymol. 2017;591:271-286. doi: 10.1016/bs.mie.2017.03.005. Epub 2017 Apr 14. Methods Enzymol. 2017. PMID: 28645372
Cited by
-
Functional regulation of the structure-specific endonuclease FEN1 by the human cytomegalovirus protein IE1 suggests a role for the re-initiation of stalled viral replication forks.PLoS Pathog. 2021 Mar 26;17(3):e1009460. doi: 10.1371/journal.ppat.1009460. eCollection 2021 Mar. PLoS Pathog. 2021. PMID: 33770148 Free PMC article.
-
Resolvases, Dissolvases, and Helicases in Homologous Recombination: Clearing the Road for Chromosome Segregation.Genes (Basel). 2020 Jan 8;11(1):71. doi: 10.3390/genes11010071. Genes (Basel). 2020. PMID: 31936378 Free PMC article. Review.
-
Recognition and processing of branched DNA substrates by Slx1-Slx4 nuclease.Nucleic Acids Res. 2019 Dec 16;47(22):11681-11690. doi: 10.1093/nar/gkz842. Nucleic Acids Res. 2019. PMID: 31584081 Free PMC article.
-
Multiple DNA repair pathways contribute to MMS-induced post-replicative DNA synthesis in S. pombe.MicroPubl Biol. 2023 Oct 2;2023:10.17912/micropub.biology.000974. doi: 10.17912/micropub.biology.000974. eCollection 2023. MicroPubl Biol. 2023. PMID: 37854101 Free PMC article.
-
CRL4Cdt2 ubiquitin ligase regulates Dna2 and Rad16 (XPF) nucleases by targeting Pxd1 for degradation.PLoS Genet. 2020 Jul 21;16(7):e1008933. doi: 10.1371/journal.pgen.1008933. eCollection 2020 Jul. PLoS Genet. 2020. PMID: 32692737 Free PMC article.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

