Contractility kits promote assembly of the mechanoresponsive cytoskeletal network
- PMID: 30559246
- PMCID: PMC6362397
- DOI: 10.1242/jcs.226704
Contractility kits promote assembly of the mechanoresponsive cytoskeletal network
Abstract
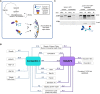
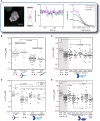
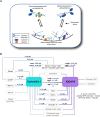
Cellular contractility is governed by a control system of proteins that integrates internal and external cues to drive diverse shape change processes. This contractility controller includes myosin II motors, actin crosslinkers and protein scaffolds, which exhibit robust and cooperative mechanoaccumulation. However, the biochemical interactions and feedback mechanisms that drive the controller remain unknown. Here, we use a proteomics approach to identify direct interactors of two key nodes of the contractility controller in the social amoeba Dictyostelium discoideum: the actin crosslinker cortexillin I and the scaffolding protein IQGAP2. We highlight several unexpected proteins that suggest feedback from metabolic and RNA-binding proteins on the contractility controller. Quantitative in vivo biochemical measurements reveal direct interactions between myosin II and cortexillin I, which form the core mechanosensor. Furthermore, IQGAP1 negatively regulates mechanoresponsiveness by competing with IQGAP2 for binding the myosin II-cortexillin I complex. These myosin II-cortexillin I-IQGAP2 complexes are pre-assembled into higher-order mechanoresponsive contractility kits (MCKs) that are poised to integrate into the cortex upon diffusional encounter coincident with mechanical inputs.This article has an associated First Person interview with the first author of the paper.
Keywords: Cortexillin I; FCCS; IQGAP; LC-MS; Myosin II; SiMPull.
© 2019. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interestsThe authors declare no competing or financial interests.
Figures


Similar articles
-
A mechanosensory system governs myosin II accumulation in dividing cells.Mol Biol Cell. 2012 Apr;23(8):1510-23. doi: 10.1091/mbc.E11-07-0601. Epub 2012 Feb 29. Mol Biol Cell. 2012. PMID: 22379107 Free PMC article.
-
Mechanical stress and network structure drive protein dynamics during cytokinesis.Curr Biol. 2015 Mar 2;25(5):663-70. doi: 10.1016/j.cub.2015.01.025. Epub 2015 Feb 19. Curr Biol. 2015. PMID: 25702575 Free PMC article.
-
Mechanosensing through cooperative interactions between myosin II and the actin crosslinker cortexillin I.Curr Biol. 2009 Sep 15;19(17):1421-8. doi: 10.1016/j.cub.2009.07.018. Epub 2009 Jul 30. Curr Biol. 2009. PMID: 19646871 Free PMC article.
-
Signaling pathways regulating Dictyostelium myosin II.J Muscle Res Cell Motil. 2002;23(7-8):703-18. doi: 10.1023/a:1024467426244. J Muscle Res Cell Motil. 2002. PMID: 12952069 Review.
-
Genetic approaches to dissect the mechanisms of two distinct pathways of cell cycle-coupled cytokinesis in Dictyostelium.Cell Struct Funct. 2001 Dec;26(6):585-91. doi: 10.1247/csf.26.585. Cell Struct Funct. 2001. PMID: 11942613 Review.
Cited by
-
Non-muscle myosin 2 filaments are processive in cells.bioRxiv [Preprint]. 2023 Feb 26:2023.02.24.529920. doi: 10.1101/2023.02.24.529920. bioRxiv. 2023. Update in: Biophys J. 2023 Sep 19;122(18):3678-3689. doi: 10.1016/j.bpj.2023.05.014. PMID: 36865321 Free PMC article. Updated. Preprint.
-
The lectin Discoidin I acts in the cytoplasm to help assemble the contractile machinery.J Cell Biol. 2022 Nov 7;221(11):e202202063. doi: 10.1083/jcb.202202063. Epub 2022 Sep 27. J Cell Biol. 2022. PMID: 36165849 Free PMC article.
-
Discovery and Quantitative Dissection of Cytokinesis Mechanisms Using Dictyostelium discoideum.Methods Mol Biol. 2024;2814:1-27. doi: 10.1007/978-1-0716-3894-1_1. Methods Mol Biol. 2024. PMID: 38954194
-
Proteomic Analysis of Dictyostelium discoideum by Mass Spectrometry.Methods Mol Biol. 2024;2814:247-255. doi: 10.1007/978-1-0716-3894-1_17. Methods Mol Biol. 2024. PMID: 38954210 Free PMC article.
-
RNA-Seq analysis of duck embryo fibroblast cells gene expression during duck Tembusu virus infection.Vet Res. 2022 May 18;53(1):34. doi: 10.1186/s13567-022-01051-y. Vet Res. 2022. PMID: 35585616 Free PMC article.
References
-
- Bai H., Zhu Q., Surcel A., Luo T., Ren Y., Guan B., Liu Y., Wu N., Joseph N. E., Wang T.-L. et al. (2016). Yes-associated protein impacts adherens junction assembly through regulating actin cytoskeleton organization. Am. J. Physiol. Gastrointest. Liver Physiol. 311, G396-G411. 10.1152/ajpgi.00027.2016 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

