Engineering RNA Virus Interference via the CRISPR/Cas13 Machinery in Arabidopsis
- PMID: 30572690
- PMCID: PMC6315463
- DOI: 10.3390/v10120732
Engineering RNA Virus Interference via the CRISPR/Cas13 Machinery in Arabidopsis
Abstract
Clustered regularly interspaced short palindromic repeats (CRISPR) and CRISPR-associated (Cas) systems are key immune mechanisms helping prokaryotic species fend off RNA and DNA viruses. CRISPR/Cas9 has broad applications in basic research and biotechnology and has been widely used across eukaryotic species for genome engineering and functional analysis of genes. The recently developed CRISPR/Cas13 systems target RNA rather than DNA and thus offer new potential for transcriptome engineering and combatting RNA viruses. Here, we used CRISPR/LshCas13a to stably engineer Arabidopsis thaliana for interference against the RNA genome of Turnip mosaic virus (TuMV). Our data demonstrate that CRISPR RNAs (crRNAs) guiding Cas13a to the sequences encoding helper component proteinase silencing suppressor (HC-Pro) or GFP target 2 (GFP-T2) provide better interference compared to crRNAs targeting other regions of the TuMV RNA genome. This work demonstrates the exciting potential of CRISPR/Cas13 to be used as an antiviral strategy to obstruct RNA viruses, and encourages the search for more robust and effective Cas13 variants or CRISPR systems that can target RNA.
Keywords: CRISPR/Cas systems; CRISPR/Cas13a; RNA interference; RNA knockdown; molecular immunity; transcriptome regulation; virus interference.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

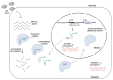
References
-
- Sudarshana M.R., Roy G., Falk B.W. Methods for engineering resistance to plant viruses. Methods Mol. Biol. 2007;354:183–195. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources

