Anti-silencing factor Epe1 associates with SAGA to regulate transcription within heterochromatin
- PMID: 30573453
- PMCID: PMC6317313
- DOI: 10.1101/gad.318030.118
Anti-silencing factor Epe1 associates with SAGA to regulate transcription within heterochromatin
Abstract
Heterochromatin is a highly condensed form of chromatin that silences gene transcription. Although high levels of transcriptional activities disrupt heterochromatin, transcription of repetitive DNA elements and subsequent processing of the transcripts by the RNAi machinery are required for heterochromatin assembly. In fission yeast, a JmjC domain protein, Epe1, promotes transcription of DNA repeats to facilitate heterochromatin formation, but overexpression of Epe1 leads to heterochromatin defects. However, the molecular function of Epe1 is not well understood. By screening the fission yeast deletion library, we found that heterochromatin defects associated with Epe1 overexpression are alleviated by mutations of the SAGA histone acetyltransferase complex. Overexpressed Epe1 associates with SAGA and recruits SAGA to heterochromatin regions, which leads to increased histone acetylation, transcription of repeats, and the disruption of heterochromatin. At its normal expression levels, Epe1 also associates with SAGA, albeit weakly. Such interaction regulates histone acetylation levels at heterochromatin and promotes transcription of repeats for heterochromatin assembly. Our results also suggest that increases of certain chromatin protein levels, which frequently occur in cancer cells, might strengthen relatively weak interactions to affect the epigenetic landscape.
Keywords: Epe1; SAGA; heterochromatin; histone; methylation; transcription.
© 2019 Bao, et al.; Published by Cold Spring Harbor Laboratory Press.
Figures




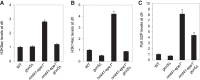
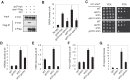
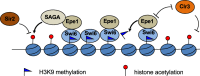
Similar articles
-
Interaction of Epe1 with the heterochromatin assembly pathway in Schizosaccharomyces pombe.Genetics. 2007 Apr;175(4):1549-60. doi: 10.1534/genetics.106.068684. Genetics. 2007. PMID: 17449867 Free PMC article.
-
The JmjC domain protein Epe1 prevents unregulated assembly and disassembly of heterochromatin.EMBO J. 2007 Nov 14;26(22):4670-82. doi: 10.1038/sj.emboj.7601892. Epub 2007 Oct 18. EMBO J. 2007. PMID: 17948055 Free PMC article.
-
Functional interaction between ELL transcription elongation factor and Epe1 reveals the role of Epe1 in the regulation of transcription outside heterochromatin.Mol Microbiol. 2021 Jul;116(1):80-96. doi: 10.1111/mmi.14691. Epub 2021 Feb 16. Mol Microbiol. 2021. PMID: 33533152
-
Destabilizing heterochromatin: Does Swi6/HP1 make the choice?Mol Cell. 2006 Jun 23;22(6):709-710. doi: 10.1016/j.molcel.2006.06.004. Mol Cell. 2006. PMID: 16793539 Review.
-
Studies on the mechanism of RNAi-dependent heterochromatin assembly.Cold Spring Harb Symp Quant Biol. 2006;71:461-71. doi: 10.1101/sqb.2006.71.044. Cold Spring Harb Symp Quant Biol. 2006. PMID: 17381328 Review.
Cited by
-
Tracking live-cell single-molecule dynamics enables measurements of heterochromatinassociated protein-protein interactions.bioRxiv [Preprint]. 2023 Oct 19:2023.03.08.531771. doi: 10.1101/2023.03.08.531771. bioRxiv. 2023. Update in: Nucleic Acids Res. 2024 Oct 14;52(18):10731-10746. doi: 10.1093/nar/gkae692. PMID: 36945633 Free PMC article. Updated. Preprint.
-
A conserved RNA degradation complex required for spreading and epigenetic inheritance of heterochromatin.Elife. 2020 Jun 3;9:e54341. doi: 10.7554/eLife.54341. Elife. 2020. PMID: 32491985 Free PMC article.
-
Nuclear Envelope Proteins Modulating the Heterochromatin Formation and Functions in Fission Yeast.Cells. 2020 Aug 16;9(8):1908. doi: 10.3390/cells9081908. Cells. 2020. PMID: 32824370 Free PMC article. Review.
-
The molecular basis of heterochromatin assembly and epigenetic inheritance.Mol Cell. 2023 Jun 1;83(11):1767-1785. doi: 10.1016/j.molcel.2023.04.020. Epub 2023 May 18. Mol Cell. 2023. PMID: 37207657 Free PMC article. Review.
-
The euchromatic histone mark H3K36me3 preserves heterochromatin through sequestration of an acetyltransferase complex in fission yeast.Microb Cell. 2020 Jan 3;7(3):80-92. doi: 10.15698/mic2020.03.711. Microb Cell. 2020. PMID: 32161768 Free PMC article.
References
-
- Abe Y, Rozqie R, Matsumura Y, Kawamura T, Nakaki R, Tsurutani Y, Tanimura-Inagaki K, Shiono A, Magoori K, Nakamura K, et al. 2015. JMJD1A is a signal-sensing scaffold that regulates acute chromatin dynamics via SWI/SNF association for thermogenesis. Nat Commun 6: 7052 10.1038/ncomms8052 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
