Interleukin 34 (IL-34) cell-surface localization regulated by the molecular chaperone 78-kDa glucose-regulated protein facilitates the differentiation of monocytic cells
- PMID: 30573681
- PMCID: PMC6378969
- DOI: 10.1074/jbc.RA118.006226
Interleukin 34 (IL-34) cell-surface localization regulated by the molecular chaperone 78-kDa glucose-regulated protein facilitates the differentiation of monocytic cells
Abstract
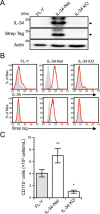
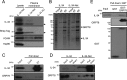
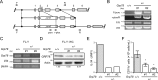
Interleukin 34 (IL-34) constitutes a cytokine that shares a common receptor, colony-stimulating factor-1 receptor (CSF-1R), with CSF-1. We recently identified a novel type of monocytic cell termed follicular dendritic cell-induced monocytic cells (FDMCs), whose differentiation depended on CSF-1R signaling through the IL-34 produced from a follicular dendritic cell line, FL-Y. Here, we report the functional mechanisms of the IL-34-mediated CSF-1R signaling underlying FDMC differentiation. CRIPSR/Cas9-mediated knockout of the Il34 gene confirmed that the ability of FL-Y cells to induce FDMCs completely depends on the IL-34 expressed by FL-Y cells. Transwell culture experiments revealed that FDMC differentiation requires a signal from a membrane-anchored form of IL-34 on the FL-Y cell surface, but not from a secreted form, in a direct interaction between FDMC precursor cells and FL-Y cells. Furthermore, flow cytometric analysis using an anti-IL-34 antibody indicated that IL-34 was also expressed on the FL-Y cell surface. Thus, we explored proteins interacting with IL-34 in FL-Y cells. Mass spectrometry analysis and pulldown assay identified that IL-34 was associated with the molecular chaperone 78-kDa glucose-regulated protein (GRP78) in the plasma membrane fraction of FL-Y cells. Consistent with this finding, GRP78-heterozygous FL-Y cells expressed a lower level of IL-34 protein on their cell surface and exhibited a reduced competency to induce FDMC differentiation compared with the original FL-Y cells. These results indicated a novel GRP78-dependent localization and specific function of IL-34 in FL-Y cells related to monocytic cell differentiation.
Keywords: CRISPR/Cas; GRP78; Western blot; cell differentiation; cell surface protein; chaperone; colony-stimulating factor-1 receptor; cytokine; follicular dendritic cell; interleukin; interleukin 34; monocyte; plasma membrane.
© 2019 Ogawa et al.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures

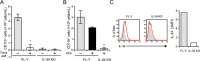
Similar articles
-
CSF-1 receptor-mediated differentiation of a new type of monocytic cell with B cell-stimulating activity: its selective dependence on IL-34.J Leukoc Biol. 2014 Jan;95(1):19-31. doi: 10.1189/jlb.0613311. Epub 2013 Sep 19. J Leukoc Biol. 2014. PMID: 24052571 Free PMC article.
-
The immunoreceptor SLAMF8 promotes the differentiation of follicular dendritic cell-dependent monocytic cells with B cell-activating ability.FEBS Lett. 2022 Oct;596(20):2659-2667. doi: 10.1002/1873-3468.14468. Epub 2022 Aug 19. FEBS Lett. 2022. PMID: 35953458
-
Expression of FLT3 receptor and response to FLT3 ligand by leukemic cells.Leukemia. 1996 Apr;10(4):588-99. Leukemia. 1996. PMID: 8618433 Review.
-
IL-21-dependent B cell death driven by prostaglandin E2, a product secreted from follicular dendritic cells.J Immunol. 2011 Oct 15;187(8):4210-8. doi: 10.4049/jimmunol.1100934. Epub 2011 Sep 12. J Immunol. 2011. PMID: 21911600
-
Interleukins and colony stimulating factors in human myeloid leukemia cell lines.Leuk Lymphoma. 1995 Dec;20(1-2):1-15. doi: 10.3109/10428199509054748. Leuk Lymphoma. 1995. PMID: 8750618 Review.
Cited by
-
The twin cytokines interleukin-34 and CSF-1: masterful conductors of macrophage homeostasis.Theranostics. 2021 Jan 1;11(4):1568-1593. doi: 10.7150/thno.50683. eCollection 2021. Theranostics. 2021. PMID: 33408768 Free PMC article. Review.
-
Loss of IL-34 Expression Indicates Poor Prognosis in Patients With Lung Adenocarcinoma.Front Oncol. 2021 Jul 16;11:639724. doi: 10.3389/fonc.2021.639724. eCollection 2021. Front Oncol. 2021. PMID: 34336646 Free PMC article.
-
The macrophage: a key player in the pathophysiology of peripheral neuropathies.J Neuroinflammation. 2022 Apr 16;19(1):97. doi: 10.1186/s12974-022-02454-6. J Neuroinflammation. 2022. PMID: 35429971 Free PMC article. Review.
-
The M-CSF receptor in osteoclasts and beyond.Exp Mol Med. 2020 Aug;52(8):1239-1254. doi: 10.1038/s12276-020-0484-z. Epub 2020 Aug 17. Exp Mol Med. 2020. PMID: 32801364 Free PMC article. Review.
References
-
- Wei S., Nandi S., Chitu V., Yeung Y.-G., Yu W., Huang M., Williams L. T., Lin H., and Stanley E. R. (2010) Functional overlap but differential expression of CSF-1 and IL-34 in their CSF-1 receptor-mediated regulation of myeloid cells. J. Leukoc. Biol. 88, 495–505 10.1189/jlb.1209822 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

