Genetic exchanges are more frequent in bacteria encoding capsules
- PMID: 30576310
- PMCID: PMC6322790
- DOI: 10.1371/journal.pgen.1007862
Genetic exchanges are more frequent in bacteria encoding capsules
Abstract
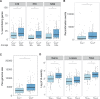
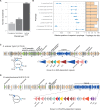
Capsules allow bacteria to colonize novel environments, to withstand numerous stresses, and to resist antibiotics. Yet, even though genetic exchanges with other cells should be adaptive under such circumstances, it has been suggested that capsules lower the rates of homologous recombination and horizontal gene transfer. We analysed over one hundred pan-genomes and thousands of bacterial genomes for the evidence of an association between genetic exchanges (or lack thereof) and the presence of a capsule system. We found that bacteria encoding capsules have larger pan-genomes, higher rates of horizontal gene transfer, and higher rates of homologous recombination in their core genomes. Accordingly, genomes encoding capsules have more plasmids, conjugative elements, transposases, prophages, and integrons. Furthermore, capsular loci are frequent in plasmids, and can be found in prophages. These results are valid for Bacteria, independently of their ability to be naturally transformable. Since we have shown previously that capsules are commonly present in nosocomial pathogens, we analysed their co-occurrence with antibiotic resistance genes. Genomes encoding capsules have more antibiotic resistance genes, especially those encoding efflux pumps, and they constitute the majority of the most worrisome nosocomial bacteria. We conclude that bacteria with capsule systems are more genetically diverse and have fast-evolving gene repertoires, which may further contribute to their success in colonizing novel niches such as humans under antibiotic therapy.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



Similar articles
-
Abundance and co-occurrence of extracellular capsules increase environmental breadth: Implications for the emergence of pathogens.PLoS Pathog. 2017 Jul 24;13(7):e1006525. doi: 10.1371/journal.ppat.1006525. eCollection 2017 Jul. PLoS Pathog. 2017. PMID: 28742161 Free PMC article.
-
Interplay between the cell envelope and mobile genetic elements shapes gene flow in populations of the nosocomial pathogen Klebsiella pneumoniae.PLoS Biol. 2021 Jul 6;19(7):e3001276. doi: 10.1371/journal.pbio.3001276. eCollection 2021 Jul. PLoS Biol. 2021. PMID: 34228700 Free PMC article.
-
Regulation of genetic flux between bacteria by restriction-modification systems.Proc Natl Acad Sci U S A. 2016 May 17;113(20):5658-63. doi: 10.1073/pnas.1603257113. Epub 2016 May 2. Proc Natl Acad Sci U S A. 2016. PMID: 27140615 Free PMC article.
-
[Homologous recombination among bacterial genomes: the measurement and identification].Yi Chuan. 2016 Feb;38(2):137-43. doi: 10.16288/j.yczz.15-382. Yi Chuan. 2016. PMID: 26907777 Review. Chinese.
-
Antibiotic resistance in Pseudomonas aeruginosa - Mechanisms, epidemiology and evolution.Drug Resist Updat. 2019 May;44:100640. doi: 10.1016/j.drup.2019.07.002. Epub 2019 Jul 19. Drug Resist Updat. 2019. PMID: 31492517 Review.
Cited by
-
ICEKp2: description of an integrative and conjugative element in Klebsiella pneumoniae, co-occurring and interacting with ICEKp1.Sci Rep. 2019 Sep 25;9(1):13892. doi: 10.1038/s41598-019-50456-x. Sci Rep. 2019. PMID: 31554924 Free PMC article.
-
Tracking and characterization of a novel conjugative transposon identified by shotgun transposon mutagenesis.Front Microbiol. 2024 Mar 26;15:1241582. doi: 10.3389/fmicb.2024.1241582. eCollection 2024. Front Microbiol. 2024. PMID: 38601936 Free PMC article.
-
Comparative genomics of Vibrio toranzoniae strains.Int Microbiol. 2025 Mar;28(3):485-496. doi: 10.1007/s10123-024-00557-z. Epub 2024 Jul 12. Int Microbiol. 2025. PMID: 38995500 Free PMC article.
-
Adaptations of Vibrio parahaemolyticus to Stress During Environmental Survival, Host Colonization, and Infection.Front Microbiol. 2021 Oct 7;12:737299. doi: 10.3389/fmicb.2021.737299. eCollection 2021. Front Microbiol. 2021. PMID: 34690978 Free PMC article. Review.
-
Polymorphism and mutational diversity of virulence (vcgCPI/vcgCPE) and resistance determinants (aac(3)-IIa, (aacC2, strA, Sul 1, and 11) among human pathogenic Vibrio species recovered from surface waters in South-Western districts of Uganda.J Genet Eng Biotechnol. 2023 Oct 6;21(1):94. doi: 10.1186/s43141-023-00554-1. J Genet Eng Biotechnol. 2023. PMID: 37801152 Free PMC article.
References
-
- Lam TT, Claus H, Frosch M, Vogel U. Sequence analysis of serotype-specific synthesis regions II of Haemophilus influenzae serotypes c and d: evidence for common ancestry of capsule synthesis in Pasteurellaceae and Neisseria meningitidis. Res Microbiol. 2011;162(5):483–7. 10.1016/j.resmic.2011.04.002 . - DOI - PubMed
-
- Mostowy RJ, Croucher NJ, De Maio N, Chewapreecha C, Salter SJ, Turner P, et al. Pneumococcal Capsule Synthesis Locus cps as Evolutionary Hotspot with Potential to Generate Novel Serotypes by Recombination. Molecular Biology and Evolution. 2017;34(10):2537–54. 10.1093/molbev/msx173 WOS:000411814800008. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

