Phosphatidylinositol 3-kinase pathway genomic alterations in 60,991 diverse solid tumors informs targeted therapy opportunities
- PMID: 30582752
- PMCID: PMC6433468
- DOI: 10.1002/cncr.31921
Phosphatidylinositol 3-kinase pathway genomic alterations in 60,991 diverse solid tumors informs targeted therapy opportunities
Abstract
Background: The phosphatidylinositol 3-kinase (PI3K) pathway is frequently altered in cancer. This report describes the landscape of PI3K alterations in solid tumors as well as co-alterations serving as potential resistance/attenuation mechanisms.
Methods: Consecutive samples were analyzed in a commercial Clinical Laboratory Improvement Amendment-certified laboratory using comprehensive genomic profiling performed by next-generation sequencing (315 genes). The co-alterations evaluated included the Erb-B2 receptor tyrosine kinase 2 (ERBB2), ERBB3, ERBB4, RAS, MET proto-oncogene tyrosine kinase (MET), and mitogen-activated protein kinase kinase (MAP2K) genes as well as tumor protein 53 (TP53), estrogen receptor 1 (ESR1), and androgen receptor (AR).
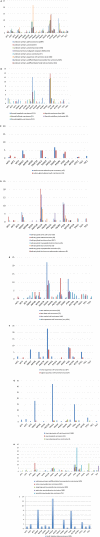
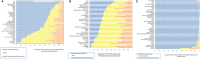
Results: Alterations in any of 18 PI3K-pathway associated genes were identified in 44% of 60,991 tumors. Although single base and insertions/deletions (indels) were the most frequent alterations, copy number changes and rearrangements were identified in 11% and 0.9% of patients, respectively. Overall, the most frequently altered genes were PIK3 catalytic subunit α (PIK3CA) (13%), phosphatase and tensin homolog (PTEN) (9%), and serine/threonine kinase 11 (STK11) (5%). Tumor types that frequently harbored at least 1 PI3K alteration were uterine (77%), cervical (62%), anal (59%), and breast (58%) cancers. Alterations also were discerned frequently in tumors with carcinosarcoma (89%) and squamous cell carcinoma (62%) histologies. Tumors with a greater likelihood of co-occurring PI3K pathway and MAPK pathway alterations included colorectal cancers (odds ratio [OR], 1.64; P < .001), mesotheliomas (OR, 2.67; P = .024), anal cancers (OR, 1.98; P = .03), and nonsquamous head and neck cancers (OR, 2.03; P = .019). The co-occurrence of ESR1 and/or AR alterations with PI3K alterations was statistically significant in bladder, colorectal, uterine, prostate, and unknown primary cancers.
Conclusions: Comprehensive genomic profiling reveals altered PI3K-related genes in 44% of solid malignancies, including rare disease and histology types. The frequency of alterations and the co-occurrence of resistance pathways vary by tumor type, directly affecting opportunities for targeted therapy.
Keywords: cancer genome; molecular profile; phosphoinositide 3-kinase catalytic subunit α (PIK3CA); precision oncology; targeted therapy.
© 2018 The Authors. Cancer published by Wiley Periodicals, Inc. on behalf of American Cancer Society.
Conflict of interest statement
Denis L. Jardim reports personal fees from Roche outside the submitted work. Razelle Kurzrock reports research funding from Incyte, Genentech, Merck, Serono, Pfizer, Sequenom, Foundation Medicine, and Guardant outside the submitted work; personal fees from X Biotech, Loxo, and Actuate Therapeutics, and Roche outside the submitted work; and has an ownership interest in Curematch Inc. Vincent A. Miller reports personal fees from Revolutions Medicines outside the submitted work. Siraj M. Ali serves on the Incysus Therapeutics Scientific Advisory Board. Sherri Z. Millis, Lee Albacher, Jeffrey S. Ross, Vincent A. Miller, and Siraj M. Ali are employees of Foundation Medicine Inc.
Figures

References
-
- Engelman JA. Targeting PI3K signaling in cancer: opportunities, challenges and limitations. Nat Rev Cancer. 2009;9:550‐562. - PubMed
-
- Millis SZ, Ikeda S, Reddy S, Gatalica Z, Kurzrock R. Landscape of phosphatidylinositol‐3‐kinase pathway alterations across 19,784 diverse solid tumors. JAMA Oncol. 2016;2:1565‐1573. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

