A Novel Transcription Factor CaSBP12 Gene Negatively Regulates the Defense Response against Phytophthora capsici in Pepper (Capsicum annuum L.)
- PMID: 30583543
- PMCID: PMC6337521
- DOI: 10.3390/ijms20010048
A Novel Transcription Factor CaSBP12 Gene Negatively Regulates the Defense Response against Phytophthora capsici in Pepper (Capsicum annuum L.)
Abstract
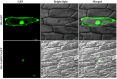
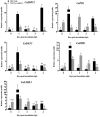
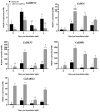
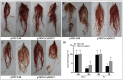
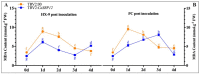
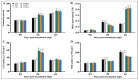
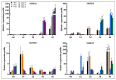
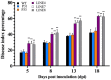
SBP-box (Squamosa-promoter binding protein) genes are a type of plant-specific transcription factor and play important roles in plant growth, signal transduction and stress response. However, little is known about the SBP-box genes in pepper (CaSBP), especially in the process of Phytophthora capsici infection. In this study, a novel gene (CaSBP12) was selected from the CaSBP gene family, which was isolated from the pepper genome database in our previous study. The CaSBP12 gene was located in the nucleus of the cell and its silencing in the pepper plant enhanced the defense response against Phytophthora capsici infection. After inoculation with Phytophthora capsici, the root activity of the CaSBP12-silenced plants is compared to control plants, while malondialdehyde (MDA) content is compared viceversa. Additionally, the expression of defense related genes (CaPO1, CaSAR8.2, CaBPR1, and CaDEF1) in the silenced plants were induced to different degrees and the peak of CaSAR8.2 and CaBPR1 were higher than that of CaDEF1. The CaSBP12 over-expressed Nicotiana benthamiana plants were more susceptible to Phytophthora capsici infection with higher EC (electrical conductivity) and MDA contents as compared to the wild-type. The relative expression of defense related genes (NbDEF, NbNPR1, NbPR1a, and NbPR1b) in transgenic and wild-type Nicotiana benthamiana plants were induced, especially the NbPR1a and NbPR1b. In conclusion, these results indicate that CaSBP12 gene negative regulates the defense response against Phytophthora capsici infection which suggests their potentially significant role in plant defense. To our knowledge, this is the first report on CaSBP gene which negative regulate defense response.
Keywords: CaSBP12; Nicotiana benthamiana; Phytophthora capsici; defense-related genes; pepper.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
CaSBP11 Participates in the Defense Response of Pepper to Phytophthora capsici through Regulating the Expression of Defense-Related Genes.Int J Mol Sci. 2020 Nov 28;21(23):9065. doi: 10.3390/ijms21239065. Int J Mol Sci. 2020. PMID: 33260627 Free PMC article.
-
Identification of Pepper CaSBP08 Gene in Defense Response Against Phytophthora capsici Infection.Front Plant Sci. 2020 Feb 26;11:183. doi: 10.3389/fpls.2020.00183. eCollection 2020. Front Plant Sci. 2020. PMID: 32174944 Free PMC article.
-
Transcription Factor CaSBP12 Negatively Regulates Salt Stress Tolerance in Pepper (Capsicum annuum L.).Int J Mol Sci. 2020 Jan 10;21(2):444. doi: 10.3390/ijms21020444. Int J Mol Sci. 2020. PMID: 31936712 Free PMC article.
-
Knockdown of the chitin-binding protein family gene CaChiIV1 increased sensitivity to Phytophthora capsici and drought stress in pepper plants.Mol Genet Genomics. 2019 Oct;294(5):1311-1326. doi: 10.1007/s00438-019-01583-7. Epub 2019 Jun 7. Mol Genet Genomics. 2019. PMID: 31175439
-
Challenges and Strategies for Breeding Resistance in Capsicum annuum to the Multifarious Pathogen, Phytophthora capsici.Front Plant Sci. 2018 May 15;9:628. doi: 10.3389/fpls.2018.00628. eCollection 2018. Front Plant Sci. 2018. PMID: 29868083 Free PMC article. Review.
Cited by
-
The CaAP2/ERF064 Regulates Dual Functions in Pepper: Plant Cell Death and Resistance to Phytophthora capsici.Genes (Basel). 2019 Jul 17;10(7):541. doi: 10.3390/genes10070541. Genes (Basel). 2019. PMID: 31319566 Free PMC article.
-
CaWRKY50 Acts as a Negative Regulator in Response to Colletotrichum scovillei Infection in Pepper.Plants (Basel). 2023 May 11;12(10):1962. doi: 10.3390/plants12101962. Plants (Basel). 2023. PMID: 37653879 Free PMC article.
-
Pepper SBP-box transcription factor, CaSBP13, plays a negatively role in drought response.Front Plant Sci. 2024 Jul 12;15:1412685. doi: 10.3389/fpls.2024.1412685. eCollection 2024. Front Plant Sci. 2024. PMID: 39070917 Free PMC article.
-
Transcriptome and metabolome analyses revealed the response mechanism of pepper roots to Phytophthora capsici infection.BMC Genomics. 2023 Oct 20;24(1):626. doi: 10.1186/s12864-023-09713-7. BMC Genomics. 2023. PMID: 37864214 Free PMC article.
-
Transcriptome analysis clarified genes involved in resistance to Phytophthora capsici in melon.PLoS One. 2020 Feb 12;15(2):e0227284. doi: 10.1371/journal.pone.0227284. eCollection 2020. PLoS One. 2020. PMID: 32050262 Free PMC article.
References
-
- Biles C.L., Brunton B.D., Wall M.M., Rivas M. Phytophthora capsici zoospore infection of pepper fruit in various physical environments. Proc. Okla. Acad. Sci. 1995;75:1–5.
-
- Mou S.L., Liu Z.Q., Gao F., Yang S., Su M.X., Shen L., Wu Y., He S.L. CaHDZ27, a Homeodomain-Leucine Zipper I (HD-Zip I) protein, positively regulates the resistance to Ralstonia solanacearum Infection in pepper. Mol. Plant-Microbe Interact. 2017;30:960–973. doi: 10.1094/MPMI-06-17-0130-R. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

