Sporocadaceae, a family of coelomycetous fungi with appendage-bearing conidia
- PMID: 30584265
- PMCID: PMC6298422
- DOI: 10.1016/j.simyco.2018.11.001
Sporocadaceae, a family of coelomycetous fungi with appendage-bearing conidia
Abstract
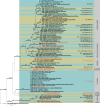
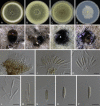
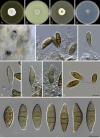
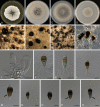
Species of Sporocadaceae are endophytic, plant pathogenic or saprobic, and associated with a wide range of host plants. Recent molecular studies that have attempted to address familial and generic boundaries of fungi belonging to Sporocadaceae were based on a limited number of samples and DNA loci. The taxonomy of this group of fungi is therefore still not fully resolved. The aim of the present study is to provide a natural classification for the Sporocadaceae based on multi-locus phylogenetic analyses, using LSU, ITS, tef-1α, tub2 and rpb2 loci, in combination with morphological data. A total of 30 well-supported monophyletic clades in Sporocadaceae are recognised, representing 23 known and seven new genera. Typifications are proposed for the type species of five genera (Diploceras, Discosia, Monochaetia, Sporocadus and Truncatella) to stabilise the application of these names. Furthermore, Neotruncatella and Dyrithiopsis are synonymised under Hymenopleella, and the generic circumscriptions of Diploceras, Disaeta, Hymenopleella, Monochaetia, Morinia, Pseudopestalotiopsis, Sarcostroma, Seimatosporium, Synnemapestaloides and Truncatella are emended. A total of 51 new species, one nomina nova and 15 combinations are introduced.
Keywords: Bartalinia pini F. Liu, L. Cai & Crous; Discosia rubi F. Liu, L. Cai & Crous; Discosia tricellularis (Okane et al.) F. Liu, L. Cai & Crous; Discosia yakushimensis (Kaz. Tanaka et al.) F. Liu, L. Cai & Crous; Distononappendiculata F. Liu, L. Cai & Crous; Distononappendiculata banksiae (Crous & Summerell) F. Liu, L. Cai & Crous; Distononappendiculata casuarinae F. Liu, L. Cai & Crous; Distononappendiculata verrucata F. Liu, L. Cai & Crous; Diversimediispora F. Liu, L. Cai & Crous; Diversimediispora humicola F. Liu, L. Cai & Crous; Heterotruncatella F. Liu, L. Cai & Crous; Heterotruncatella acacigena F. Liu, L. Cai & Crous; Heterotruncatella aspera F. Liu, L. Cai & Crous; Heterotruncatella avellanea F. Liu, L. Cai & Crous; Heterotruncatella breviappendiculata F. Liu, L. Cai & Crous; Heterotruncatella constricta F. Liu, L. Cai & Crous; Heterotruncatella diversa F. Liu, L. Cai & Crous; Heterotruncatella grevilleae F. Liu, L. Cai & Crous; Heterotruncatella longissima F. Liu, L. Cai & Crous; Heterotruncatella lutea (H.J. Swart & D.A. Griffiths) F. Liu, L. Cai & Crous; Heterotruncatella proteicola F. Liu, L. Cai & Crous; Heterotruncatella quercicola F. Liu, L. Cai & Crous; Heterotruncatella restionacearum (S.J. Lee & Crous) F. Liu, L. Cai & Crous; Heterotruncatella singularis F. Liu, L. Cai & Crous; Heterotruncatella spadicea (S.J. Lee & Crous) F. Liu, L. Cai & Crous; Heterotruncatella spartii (Senan. et al.) F. Liu, L. Cai & Crous; Heterotruncatella synapheae F. Liu, L. Cai & Crous; Heterotruncatella vinaceobubalina F. Liu, L. Cai & Crous; Hymenopleella austroafricana F. Liu, L. Cai & Crous; Hymenopleella endophytica (Hyang B. Lee et al.) F. Liu, L. Cai & Crous; Hymenopleella lakefuxianensis (L. Cai et al.) F. Liu, L. Cai & Crous; Hymenopleella polyseptata F. Liu, L. Cai & Crous; Hymenopleella subcylindrica F. Liu, L. Cai & Crous; Monochaetia quercus F. Liu, L. Cai & Crous; Morinia acaciae (Crous) F. Liu, L. Cai & Crous; Morinia crini F. Liu, L. Cai & Crous; Multi-locus phylogeny; New taxa; Nonappendiculata F. Liu, L. Cai & Crous; Nonappendiculata quercina F. Liu, L. Cai & Crous; Parabartalinia F. Liu, L. Cai & Crous; Parabartalinia lateralis F. Liu, L. Cai & Crous; Pestalotia hypericina Ces.; Pestalotia monochaeta Desmazières; Pestalotiopsis hispanica F. Liu, L. Cai & Crous; Pestalotiopsis leucadendri F. Liu, L. Cai & Crous; Pestalotiopsis spathuliappendiculata F. Liu, L. Cai & Crous; Pestalotiopsis terricola F. Liu, L. Cai & Crous; Pseudopestalotiopsis elaeidis (C. Booth & J.S. Robertson) F. Liu, L. Cai & Crous; Pseudopestalotiopsis solicola F. Liu, L. Cai & Crous; Pseudosarcostroma F. Liu, L. Cai & Crous; Pseudosarcostroma osyridicola F. Liu, L. Cai & Crous; Robillarda australiana F. Liu, L. Cai & Crous; Sarcostroma africanum F. Liu, L. Cai & Crous; Sarcostroma australiense F. Liu, L. Cai & Crous; Sarcostroma diversiseptatum F. Liu, L. Cai & Crous; Sarcostroma leucospermi F. Liu, L. Cai & Crous; Sarcostroma longiappendiculatum F. Liu, L. Cai & Crous; Sarcostroma paragrevilleae F. Liu, L. Cai & Crous; Sarcostroma proteae F. Liu, L. Cai & Crous; Seimatosporium; Seimatosporium germanicum F. Liu, L. Cai & Crous; Seimatosporium soli F. Liu, L. Cai & Crous; Seimatosporium vitis-viniferae F. Liu, L. Cai & Crous; Sphaeria artocreas Tode; Sporocadus; Sporocadus biseptatus F. Liu, L. Cai & Crous; Sporocadus cornicola (Wijayaw. & Camporesi) F. Liu, L. Cai & Crous; Sporocadus cotini F. Liu, L. Cai & Crous; Sporocadus incanus F. Liu, L. Cai & Crous; Sporocadus lichenicola Corda; Sporocadus mali F. Liu, L. Cai & Crous; Sporocadus microcyclus F. Liu, L. Cai & Crous; Sporocadus multiseptatus F. Liu, L. Cai & Crous; Sporocadus rosarum (Henn.) F. Liu, L. Cai & Crous; Sporocadus rosigena F. Liu, L. Cai & Crous; Sporocadus rotundatus F. Liu, L. Cai & Crous; Sporocadus sorbi (Wijayaw. et al.) F. Liu, L. Cai & Crous; Sporocadus trimorphus F. Liu, L. Cai & Crous; Stilbospora angustata Pers; Synnemapestaloides juniperi F. Liu, L. Cai & Crous; Taxonomy; Truncatella spadicea S. Lee & Crous; Xenoseimatosporium F. Liu, L. Cai & Crous; Xenoseimatosporium quercinum (Goonas. et al.) F. Liu, L. Cai & Crous.
Figures




















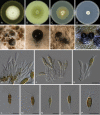
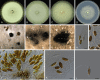


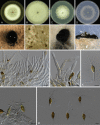
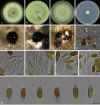
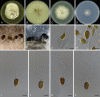

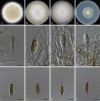
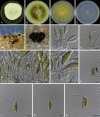


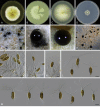









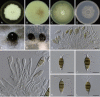
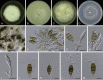
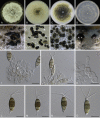


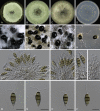






















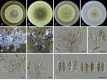


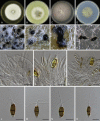

Similar articles
-
Sporocadaceae revisited.Stud Mycol. 2024 Dec;109:155-272. doi: 10.3114/sim.2024.109.03. Epub 2024 Jun 27. Stud Mycol. 2024. PMID: 39717655 Free PMC article.
-
Redisposition of acremonium-like fungi in Hypocreales.Stud Mycol. 2023 Jun;105:23-203. doi: 10.3114/sim.2023.105.02. Epub 2023 Jun 2. Stud Mycol. 2023. PMID: 38895703 Free PMC article.
-
Resolving the Phoma enigma.Stud Mycol. 2015 Sep;82:137-217. doi: 10.1016/j.simyco.2015.10.003. Epub 2015 Nov 26. Stud Mycol. 2015. PMID: 26955202 Free PMC article.
-
Fusarium: more than a node or a foot-shaped basal cell.Stud Mycol. 2021 Aug 17;98:100116. doi: 10.1016/j.simyco.2021.100116. eCollection 2021 Apr. Stud Mycol. 2021. PMID: 34466168 Free PMC article.
-
The phoma-like dilemma.Stud Mycol. 2020 May 21;96:309-396. doi: 10.1016/j.simyco.2020.05.001. eCollection 2020 Jun. Stud Mycol. 2020. PMID: 32904212 Free PMC article.
Cited by
-
A Novel Seimatosporium and Other Sporocadaceae Species Associated with Grapevine Trunk Diseases in Cyprus.Plants (Basel). 2022 Oct 16;11(20):2733. doi: 10.3390/plants11202733. Plants (Basel). 2022. PMID: 36297757 Free PMC article.
-
Molecular Phylogeny and Morphology of Amphisphaeria (= Lepteutypa) (Amphisphaeriaceae).J Fungi (Basel). 2020 Sep 17;6(3):174. doi: 10.3390/jof6030174. J Fungi (Basel). 2020. PMID: 32957501 Free PMC article.
-
Morpho-molecular characterisation of Arecophila, with A.australis and A.clypeata sp. nov. and A.miscanthi comb. nov.MycoKeys. 2022 Apr 13;88:123-149. doi: 10.3897/mycokeys.88.79475. eCollection 2022. MycoKeys. 2022. PMID: 35585934 Free PMC article.
-
Sporocadaceae revisited.Stud Mycol. 2024 Dec;109:155-272. doi: 10.3114/sim.2024.109.03. Epub 2024 Jun 27. Stud Mycol. 2024. PMID: 39717655 Free PMC article.
-
Fungal species associated with grapevine trunk diseases in Washington wine grapes and California table grapes, with novelties in the genera Cadophora, Cytospora, and Sporocadus.Front Fungal Biol. 2022 Oct 7;3:1018140. doi: 10.3389/ffunb.2022.1018140. eCollection 2022. Front Fungal Biol. 2022. PMID: 37746176 Free PMC article.
References
-
- Ariyawansa H.A., Hyde K.D., Jayasiri S.C. Fungal diversity notes 111–252—taxonomic and phylogenetic contributions to fungal taxa. Fungal Diversity. 2015;75:27–274.
-
- Batista A.C., Lima JA de, Vasconcelos CT de. Fungos exóticos achados na atmosfera do Recife. Publicações do Instituto de Micologia da Universidade do Recife. 1960;276:1–18.
-
- Bonar L. Studies on some California fungi. Mycologia. 1928;20:292–300.
LinkOut - more resources
Full Text Sources
