Coupled monoubiquitylation of the co-E3 ligase DCNL1 by Ariadne-RBR E3 ubiquitin ligases promotes cullin-RING ligase complex remodeling
- PMID: 30587576
- PMCID: PMC6393609
- DOI: 10.1074/jbc.RA118.005861
Coupled monoubiquitylation of the co-E3 ligase DCNL1 by Ariadne-RBR E3 ubiquitin ligases promotes cullin-RING ligase complex remodeling
Abstract
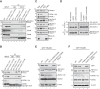
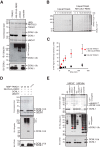
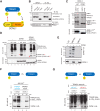
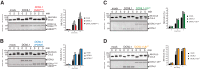
Cullin-RING E3 ubiquitin ligases (CRLs) are large and diverse multisubunit protein complexes that contribute to about one-fifth of ubiquitin-dependent protein turnover in cells. CRLs are activated by the attachment of the ubiquitin-like protein neural precursor cell expressed, developmentally down-regulated 8 (NEDD8) to the cullin subunits. This cullin neddylation is essential for a plethora of CRL-regulated cellular processes and is vital for life. In mammals, neddylation is promoted by the five co-E3 ligases, defective in cullin neddylation 1 domain-containing 1-5 (DCNL1-5); however, their functional regulation within the CRL complex remains elusive. We found here that the ubiquitin-associated (UBA) domain-containing DCNL1 is monoubiquitylated when bound to CRLs and that this monoubiquitylation depends on the CRL-associated Ariadne RBR ligases TRIAD1 (ARIH2) and HHARI (ARIH1) and strictly requires the DCNL1's UBA domain. Reconstitution of DCNL1 monoubiquitylation in vitro revealed that autoubiquitylated TRIAD1 mediates binding to the UBA domain and subsequently promotes a single ubiquitin attachment to DCNL1 in a mechanism previously dubbed coupled monoubiquitylation. Moreover, we provide evidence that DCNL1 monoubiquitylation is required for efficient CRL activity, most likely by remodeling CRLs and their substrate receptors. Collectively, this work identifies DCNL1 as a critical target of Ariadne RBR ligases and coupled monoubiquitylation of DCNL1 as an integrated mechanism that affects CRL activity and client-substrate ubiquitylation at multiple levels.
Keywords: post-translational modification; ubiquitin; ubiquitin ligase; ubiquitylation (ubiquitination); cell signaling; ARIH1; ARIH2; coupled monoubiquitylation; DCNL1; NEDD8 E3 ligase.
© 2019 Kelsall et al.
Conflict of interest statement
This work was supported by RCUK, Medical Research Council, and Max-Planck-Gesellschaft. The authors declare that they have no conflicts of interest with the contents of this article
Figures

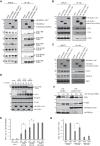
Similar articles
-
TRIAD1 and HHARI bind to and are activated by distinct neddylated Cullin-RING ligase complexes.EMBO J. 2013 Oct 30;32(21):2848-60. doi: 10.1038/emboj.2013.209. Epub 2013 Sep 27. EMBO J. 2013. PMID: 24076655 Free PMC article.
-
Two Distinct Types of E3 Ligases Work in Unison to Regulate Substrate Ubiquitylation.Cell. 2016 Aug 25;166(5):1198-1214.e24. doi: 10.1016/j.cell.2016.07.027. Cell. 2016. PMID: 27565346 Free PMC article.
-
Modulation of Cullin-RING E3 ubiquitin ligase-dependent ubiquitination by small molecule compounds.J Biol Chem. 2024 Mar;300(3):105752. doi: 10.1016/j.jbc.2024.105752. Epub 2024 Feb 13. J Biol Chem. 2024. PMID: 38354780 Free PMC article.
-
Structural Biology of CRL Ubiquitin Ligases.Adv Exp Med Biol. 2020;1217:9-31. doi: 10.1007/978-981-15-1025-0_2. Adv Exp Med Biol. 2020. PMID: 31898219 Review.
-
NEDD8-its role in the regulation of Cullin-RING ligases.Curr Opin Plant Biol. 2018 Oct;45(Pt A):112-119. doi: 10.1016/j.pbi.2018.05.017. Epub 2018 Jun 15. Curr Opin Plant Biol. 2018. PMID: 29909289 Review.
Cited by
-
The ubiquitin ligation machinery in the defense against bacterial pathogens.EMBO Rep. 2021 Nov 4;22(11):e52864. doi: 10.15252/embr.202152864. Epub 2021 Sep 13. EMBO Rep. 2021. PMID: 34515402 Free PMC article. Review.
-
Co-adaptor driven assembly of a CUL3 E3 ligase complex.Mol Cell. 2022 Feb 3;82(3):585-597.e11. doi: 10.1016/j.molcel.2022.01.004. Mol Cell. 2022. PMID: 35120648 Free PMC article.
-
The roles of E3 ligases in Hepatocellular carcinoma.Am J Cancer Res. 2022 Mar 15;12(3):1179-1214. eCollection 2022. Am J Cancer Res. 2022. PMID: 35411231 Free PMC article. Review.
-
Functional roles of E3 ubiquitin ligases in gastric cancer.Oncol Lett. 2020 Oct;20(4):22. doi: 10.3892/ol.2020.11883. Epub 2020 Jul 16. Oncol Lett. 2020. PMID: 32774495 Free PMC article. Review.
-
Chain reactions: molecular mechanisms of RBR ubiquitin ligases.Biochem Soc Trans. 2020 Aug 28;48(4):1737-1750. doi: 10.1042/BST20200237. Biochem Soc Trans. 2020. PMID: 32677670 Free PMC article. Review.
References
-
- Soucy T. A., Smith P. G., Milhollen M. A., Berger A. J., Gavin J. M., Adhikari S., Brownell J. E., Burke K. E., Cardin D. P., Critchley S., Cullis C. A., Doucette A., Garnsey J. J., Gaulin J. L., Gershman R. E., et al. (2009) An inhibitor of NEDD8-activating enzyme as a new approach to treat cancer. Nature 458, 732–736 10.1038/nature07884 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

