The structure of the SAM/SAH-binding riboswitch
- PMID: 30590743
- PMCID: PMC6411933
- DOI: 10.1093/nar/gky1283
The structure of the SAM/SAH-binding riboswitch
Abstract
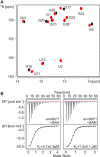
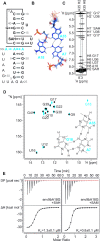
S-adenosylmethionine (SAM) is a central metabolite since it is used as a methyl group donor in many different biochemical reactions. Many bacteria control intracellular SAM concentrations using riboswitch-based mechanisms. A number of structurally different riboswitch families specifically bind to SAM and mainly regulate the transcription or the translation of SAM-biosynthetic enzymes. In addition, a highly specific riboswitch class recognizes S-adenosylhomocysteine (SAH)-the product of SAM-dependent methyl group transfer reactions-and regulates enzymes responsible for SAH hydrolysis. High-resolution structures are available for many of these riboswitch classes and illustrate how they discriminate between the two structurally similar ligands SAM and SAH. The so-called SAM/SAH riboswitch class binds both ligands with similar affinities and is structurally not yet characterized. Here, we present a high-resolution nuclear magnetic resonance structure of a member of the SAM/SAH-riboswitch class in complex with SAH. Ligand binding induces pseudoknot formation and sequestration of the ribosome binding site. Thus, the SAM/SAH-riboswitches are translational 'OFF'-switches. Our results establish a structural basis for the unusual bispecificity of this riboswitch class. In conjunction with genomic data our structure suggests that the SAM/SAH-riboswitches might be an evolutionary late invention and not a remnant of a primordial RNA-world as suggested for other riboswitches.
© The Author(s) 2018. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures





References
-
- Fontecave M., Atta M., Mulliez E.. S-adenosylmethionine. Nothing goes to waste. Trends Biochem. Sci. 2004; 29:243–249. - PubMed
-
- Struck A.-W., Thompson M.L., Wong L.S., Micklefield J.. S-adenosyl-methionine-dependent methyltransferases. Highly versatile enzymes in biocatalysis, biosynthesis and other biotechnological applications. Chembiochem. 2012; 13:2642–2655. - PubMed
-
- Lee J.E., Cornell K.A., Riscoe M.K., Howell P.L.. Structure of Escherichia coli 5′-methylthioadenosine/ S-adenosylhomocysteine nucleosidase inhibitor complexes provide insight into the conformational changes required for substrate binding and catalysis. J. Biol. Chem. 2003; 278:8761–8770. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

