Immune checkpoint inhibitors in MITF family translocation renal cell carcinomas and genetic correlates of exceptional responders
- PMID: 30591082
- PMCID: PMC6307255
- DOI: 10.1186/s40425-018-0482-z
Immune checkpoint inhibitors in MITF family translocation renal cell carcinomas and genetic correlates of exceptional responders
Abstract
Background: Microphthalmia Transcription Factor (MITF)family translocation renal cell carcinoma (tRCC) is a rare RCC subtype harboring TFE3/TFEB translocations. The prognosis in the metastatic (m) setting is poor. Programmed death ligand-1 expression was reported in 90% of cases, prompting us to analyze the benefit of immune checkpoint inhibitors (ICI) in this population.
Patients and methods: This multicenter retrospective study identified patients with MITF family mtRCC who had received an ICI in any of 12 referral centers in France or the USA. Response rate according to RECIST criteria, progression-free survival (PFS), and overall survival (OS) were analyzed. Genomic alterations associated with response were determined for 8 patients.
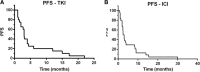
Results: Overall, 24 patients with metastatic disease who received an ICI as second or later line of treatment were identified. Nineteen (82.6%) of these patients had received a VEGFR inhibitor as first-line treatment, with a median PFS of 3 months (range, 1-22 months). The median PFS for patients during first ICI treatment was 2.5 months (range, 1-40 months); 4 patients experienced partial response (16,7%) and 3 (12,5%) had stable disease. Of the patients whose genomic alterations were analyzed, two patients with mutations in bromodomain-containing genes (PBRM1 and BRD8) had a clinical benefit. Resistant clones in a patient with exceptional response to ipilimumab showed loss of BRD8 mutations and increased mutational load driven by parallel evolution affecting 17 genes (median mutations per gene, 3), which were enriched mainly for O-glycan processing (29.4%, FDR = 9.7 × 10- 6).
Conclusions: MITF family tRCC is an aggressive disease with similar responses to ICIs as clear-cell RCC. Mutations in bromodomain-containing genes might be associated with clinical benefit. The unexpected observation about parallel evolution of genes involved in O-glycosylation as a mechanism of resistance to ICI warrants exploration.
Keywords: Antiangiogenic agents; Bromodomain-containing genes; O-glycosylation; Parallel evolution; TFE3; TFEB.
Conflict of interest statement
Ethics approval and consent to participate
Patient data were collected in compliance with the IRB guidelines of each participating institution. All study protocols were performed in accordance with the ethical tenets of the Declaration of Helsinki.
Consent for publication
Written informed consent was obtained from all patients for whom genomic testing was performed. Oral informed consent was obtained from all other patients.
Competing interests
Dr. Malouf reports consultancy fees from BMS, Pfizer, Ibsen, Novartis, Astellas, and grants from Novartis and Pfizer, outside the submitted work. Dr. Barthélémy reports consultancy fees from BMS, Pfizer, Novartis, outside the submitted work. Dr. Chevreau reports consultancy fees from BMS, Ipsen, Pfizer, Novartis,outside the submitted work. Dr. Borchiellini reports consulting fees from Bristol-Myers Squibb, Ipsen, Novartis, Pfizer, outside the submitted work. Dr. Voss reports consulting fees from Exelixis, Novartis, Eisai, Calithera, Alexion, grants from Genentech, BMS, outside the submitted work. Dr. George reports consultancy fees from Bristol-Myers Squibb, Novartis, Bayer, Pfizer, Exelixis, AstraZeneca, Janssen Oncology, Corvus Pharmaceuticals, Genentech/Roche, and research grants from Pfizer, Acceleron Pharma, Merck, Agensys, Novartis, Bristol-Myers Squibb, Bayer. Dr. Chism reports consultancy fees from Karyopharm Inc., Pfizer, Exelixis outside the submitted work. Dr. Tannir reports consulting fees from BMS, Exelixis, Nektar, Novartis, Pfizer, Argos, Calithera and grants from BMS, Exelixis, Epizyme, Novartis, Miranti, outside the submitted work, outside the submitted work. Dr. Choueiri has reports 1) research grants (Institutional and personal) from AstraZeneca, Bayer, BMS, Cerulean, Eisai, Foundation Medicine Inc., Exelixis, Ipsen, Tracon, Genentech, Roche, Roche Products Limited, GlaxoSmithKline, Merck, Novartis, Peloton, Pfizer, Prometheus Labs, Corvus, Calithera, Analysis Group, Takeda. 2) Honoraria: AstraZeneca, Alexion, Sanofi/Aventis, Bayer, BMS, Cerulean, Eisai, Foundation Medicine Inc., Exelixis, Genentech, Roche, GlaxoSmithKline, Merck, Novartis, Peloton, Pfizer, EMD Serono, Prometheus Labs, Corvus, Ipsen, Up-to-Date, NCCN, Analysis Group, NCCN, Michael J. Hennessy (MJH) Associates, Inc (Healthcare Communications Company with several brands such as OnClive and PER), L-path, Kidney Cancer Journal, Clinical Care Options, Platform Q, Navinata Healthcare, Harborside Press, American Society of Medical Oncology, NEJM, Lancet Oncology, Heron Therapeutics. • 3) Consulting or Advisory Role: AstraZeneca, Alexion, Sanofi/Aventis, Bayer, BMS, Cerulean, Eisai, Foundation Medicine Inc., Exelixis, Genentech, Heron Therapeutics, Roche, GlaxoSmithKline, Merck, Novartis, Peloton, Pfizer, EMD Serono, Prometheus Labs, Corvus, Ipsen, Up-to-Date, NCCN, Analysis Group. • No speaker’s bureau • No leadership or employment in for-profit companies (other present or past leadership roles: Director of GU Oncology Division at Dana-Farber and past President of medical Staff at Dana-Farber), member of NCCN Kidney panel and the GU Steering Committee, past chairman of the Kidney cancer Association Medical and Scientific Steering Committee) • No Patents, royalties or other intellectual properties • Travel, accommodations, expenses, in relation to consulting, advisory roles, or honoraria • Medical writing and editorial assistance support may have been funded by Communications companies funded by pharmaceutical companies • The institution (Dana-Farber Cancer Institute) may have received additional independent funding of drug companies or/and royalties potentially involved in research around the subject matter.Dr. Boilève, Dr. Carlo, Dr. Oudard, Dr. Landman-Parker, Dr. Tabone, Dr. Amin, Dr. M. A Bilen, Dr. Bosse, Dr. A. Coulomb-L’hermine and Dr. Su have nothing to disclose.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures



References
-
- Delahunt B, Srigley JR, Egevad L, et al. International Society of Urological Pathology grading and other prognostic factors for renal neoplasia. Eur Urol. 2014;66(5). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous
