PAX3-FOXO1 escapes miR-495 regulation during muscle differentiation
- PMID: 30593263
- PMCID: PMC6380322
- DOI: 10.1080/15476286.2018.1564464
PAX3-FOXO1 escapes miR-495 regulation during muscle differentiation
Abstract
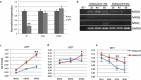
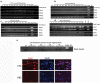
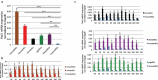
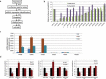
Pax3 plays an essential role in myogenesis. Previously, we found a tumor-signature chimeric fusion RNA, PAX3-FOXO1 also present during muscle differentiation, raising the possibility of its physiological role. Here we demonstrated that the fusion is needed transiently for muscle lineage commitment. Interestingly, the fusion ortholog was not found in seven mouse muscle differentiation/regeneration systems, nor in other stem cell differentiation systems of another three mammal species. We noticed that Pax3 is expressed at a much lower level in human stem cells, and during muscle differentiation than in other mammals. Given the fact that the fusion and the parental Pax3 share common downstream targets, we reasoned that forming the fusion may be a mechanism for human cells to escape certain microRNA regulation on Pax3. By sequence comparison, we identified 16 candidate microRNAs that may specifically target the human PAX3 3'UTR. We used a luciferase reporter assay, examined the microRNAs expression, and conducted mutagenesis on the reporters, as well as a CRISPR/Cas9 mediated editing on the endogenous allele. Finally, we identified miR-495 as a microRNA that specifically targets human PAX3. Examining several other fusion RNAs revealed that the human-specificity is not limited to PAX3-FOXO1. Based on these observations, we conclude that PAX3-FOXO1 fusion RNA is absent in mouse, or other mammals we tested, the fusion RNA is a mechanism to escape microRNA, miR-495 regulation in humans, and that it is not the only human-specific fusion RNA.
Keywords: Chimeric RNA; adipose-derived stroma/stem cells; mesenchymal stem cells; miR-495; microRNA; myogenesis.
Figures


Similar articles
-
PAX3-FOXO1 transgenic zebrafish models identify HES3 as a mediator of rhabdomyosarcoma tumorigenesis.Elife. 2018 Jun 5;7:e33800. doi: 10.7554/eLife.33800. Elife. 2018. PMID: 29869612 Free PMC article.
-
Acquisition of an oncogenic fusion protein serves as an initial driving mutation by inducing aneuploidy and overriding proliferative defects.Oncotarget. 2016 Sep 27;7(39):62814-62835. doi: 10.18632/oncotarget.11716. Oncotarget. 2016. PMID: 27588498 Free PMC article.
-
Fusion transcriptome profiling provides insights into alveolar rhabdomyosarcoma.Proc Natl Acad Sci U S A. 2016 Nov 15;113(46):13126-13131. doi: 10.1073/pnas.1612734113. Epub 2016 Oct 31. Proc Natl Acad Sci U S A. 2016. PMID: 27799565 Free PMC article.
-
PAX3-FOXO1: Zooming in on an "undruggable" target.Semin Cancer Biol. 2018 Jun;50:115-123. doi: 10.1016/j.semcancer.2017.11.006. Epub 2017 Nov 14. Semin Cancer Biol. 2018. PMID: 29146205 Review.
-
Post-transcriptional regulation of myogenic transcription factors during muscle development and pathogenesis.J Muscle Res Cell Motil. 2024 Mar;45(1):21-39. doi: 10.1007/s10974-023-09663-3. Epub 2024 Jan 11. J Muscle Res Cell Motil. 2024. PMID: 38206489 Review.
Cited by
-
RNA fusion in human retinal development.Elife. 2024 Jan 2;13:e92523. doi: 10.7554/eLife.92523. Elife. 2024. PMID: 38165397 Free PMC article.
-
Functional heritage: the evolution of chimeric RNA into a gene.RNA Biol. 2020 Jan;17(1):125-134. doi: 10.1080/15476286.2019.1670038. Epub 2019 Sep 29. RNA Biol. 2020. PMID: 31566065 Free PMC article.
-
The landscape of chimeric RNAs in non-diseased tissues and cells.Nucleic Acids Res. 2020 Feb 28;48(4):1764-1778. doi: 10.1093/nar/gkz1223. Nucleic Acids Res. 2020. PMID: 31965184 Free PMC article.
-
NLRP3 Inflammasome Activation by MicroRNA-495 Promoter Methylation May Contribute to the Progression of Acute Lung Injury.Mol Ther Nucleic Acids. 2019 Dec 6;18:801-814. doi: 10.1016/j.omtn.2019.08.028. Epub 2019 Oct 3. Mol Ther Nucleic Acids. 2019. PMID: 31734560 Free PMC article.
-
Chimeric RNA RRM2-C2orf48 plays an oncogenic role in the development of NNK-induced lung cancer.iScience. 2022 Dec 2;26(1):105708. doi: 10.1016/j.isci.2022.105708. eCollection 2023 Jan 20. iScience. 2022. PMID: 36570773 Free PMC article.
References
-
- Yuan H, Qin F, Movassagh M, et al. A chimeric RNA characteristic of rhabdomyosarcoma in normal myogenesis process. Cancer Discov. 2013;3:1394–1403. - PubMed
-
- Ridgeway AG, Skerjanc IS.. Pax3 is essential for skeletal myogenesis and the expression of Six1 and Eya2. J Biol Chem. 2001;276:19033–19039. - PubMed
-
- Bennicelli JL, Advani S, Schafer BW, et al. PAX3 and PAX7 exhibit conserved cis-acting transcription repression domains and utilize a common gain of function mechanism in alveolar rhabdomyosarcoma. Oncogene. 1999;18:4348–4356. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
