Characterization of the mitochondrial genome of Arge bella Wei & Du sp. nov. (Hymenoptera: Argidae)
- PMID: 30595984
- PMCID: PMC6305119
- DOI: 10.7717/peerj.6131
Characterization of the mitochondrial genome of Arge bella Wei & Du sp. nov. (Hymenoptera: Argidae)
Abstract
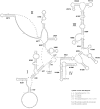
We describe Arge bella Wei & Du sp. nov., a large and beautiful species of Argidae from south China, and report its mitochondrial genome based on high-throughput sequencing data. We present the gene order, nucleotide composition of protein-coding genes (PCGs), and the secondary structures of RNA genes. The nearly complete mitochondrial genome of A. bella has a length of 15,576 bp and a typical set of 37 genes (22 tRNAs, 13 PCGs, and 2 rRNAs). Three tRNAs are rearranged in the A. bella mitochondrial genome as compared to the ancestral type in insects: trnM and trnQ are shuffled, while trnW is translocated from the trnW-trnC-trnY cluster to a location downstream of trnI. All PCGs are initiated by ATN codons, and terminated with TAA, TA or T as stop codons. All tRNAs have a typical cloverleaf secondary structure, except for trnS1. H821 of rrnS and H976 of rrnL are redundant. A phylogenetic analysis based on mitochondrial genome sequences of A. bella, 21 other symphytan species, two apocritan representatives, and four outgroup taxa supports the placement of Argidae as sister to the Pergidae within the symphytan superfamily Tenthredinoidea.
Keywords: Arge bella; Cladistic analysis; Mitochondrial genome; New species; Secondary structure; Taxonomy.
Conflict of interest statement
The authors declare there are no competing interests.
Figures






References
-
- Abe M, Smith DR. The genus-group names of Symphyta (Hymenoptera) and their type species. Esakia, Fukuoka. 1991;31:1–115.
-
- Benson RB. On the classification of sawflies (Hymenoptera Symphyta) The Transactions of the Royal Entomological Society of London. 1938;87(15):353–384.
-
- Cannone JJ, Subramanian S, Schnare MN, Collett JR, D’Souza LM, Du Y, Feng B, Lin N, Madabusi LV, Müller KM, Pande N, Shang Z, Yu N, Gutell RR. The comparative RNA web (CRW) site: an online database of comparative sequence and structure information for ribosomal, intron, and other RNAs. BMC Bioinformatics. 2002;3:1–31. doi: 10.1186/1471-2105-3-2. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

