p53 reactivation with induction of massive apoptosis-1 (PRIMA-1) inhibits amyloid aggregation of mutant p53 in cancer cells
- PMID: 30602570
- PMCID: PMC6416452
- DOI: 10.1074/jbc.RA118.004671
p53 reactivation with induction of massive apoptosis-1 (PRIMA-1) inhibits amyloid aggregation of mutant p53 in cancer cells
Abstract
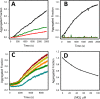
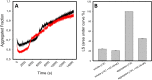
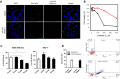
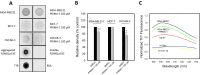
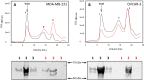
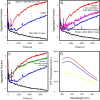
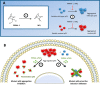
p53 mutants can form amyloid-like structures that accumulate in cells. p53 reactivation with induction of massive apoptosis-1 (PRIMA-1) and its primary active metabolite, 2-methylene-3-quinuclidinone (MQ), can restore unfolded p53 mutants to a native conformation that induces apoptosis and activates several p53 target genes. However, whether PRIMA-1 can clear p53 aggregates is unclear. In this study, we investigated whether PRIMA-1 can restore aggregated mutant p53 to a native form. We observed that the p53 mutant protein is more sensitive to both PRIMA-1 and MQ aggregation inhibition than WT p53. The results of anti-amyloid oligomer antibody assays revealed that PRIMA-1 reverses mutant p53 aggregate accumulation in cancer cells. Size-exclusion chromatography of the lysates from mutant p53-containing breast cancer and ovarian cell lines confirmed that PRIMA-1 substantially decreases p53 aggregates. We also show that MDA-MB-231 cell lysates can "seed" aggregation of the central core domain of recombinant WT p53, corroborating the prion-like behavior of mutant p53. We also noted that this aggregation effect was inhibited by MQ and PRIMA-1. This study provides the first demonstration that PRIMA-1 can rescue amyloid-state p53 mutants, a strategy that could be further explored as a cancer treatment.
Keywords: amyloid; anticancer drug; p53; protein aggregation; protein misfolding.
© 2019 Rangel et al.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures


Similar articles
-
Structural basis of reactivation of oncogenic p53 mutants by a small molecule: methylene quinuclidinone (MQ).Nat Commun. 2021 Dec 3;12(1):7057. doi: 10.1038/s41467-021-27142-6. Nat Commun. 2021. PMID: 34862374 Free PMC article.
-
PRIMA-1 inhibits growth of breast cancer cells by re-activating mutant p53 protein.Int J Oncol. 2009 Nov;35(5):1015-23. doi: 10.3892/ijo_00000416. Int J Oncol. 2009. PMID: 19787255
-
Proteomic identification of heat shock protein 90 as a candidate target for p53 mutation reactivation by PRIMA-1 in breast cancer cells.Breast Cancer Res. 2005;7(5):R765-74. doi: 10.1186/bcr1290. Epub 2005 Jul 27. Breast Cancer Res. 2005. PMID: 16168122 Free PMC article.
-
Prima-1 and APR-246 in Cancer Therapy.Klin Onkol. 2018 Winter;31(Suppl 2):71-76. doi: 10.14735/amko20182S71. Klin Onkol. 2018. PMID: 31023027 Review. English.
-
Prion-like aggregation of mutant p53 in cancer.Trends Biochem Sci. 2014 Jun;39(6):260-7. doi: 10.1016/j.tibs.2014.04.001. Epub 2014 Apr 26. Trends Biochem Sci. 2014. PMID: 24775734 Review.
Cited by
-
Protein of a thousand faces: The tumor-suppressive and oncogenic responses of p53.Front Mol Biosci. 2022 Aug 25;9:944955. doi: 10.3389/fmolb.2022.944955. eCollection 2022. Front Mol Biosci. 2022. PMID: 36090037 Free PMC article. Review.
-
Sulfated glycosaminoglycans mediate prion-like behavior of p53 aggregates.Proc Natl Acad Sci U S A. 2020 Dec 29;117(52):33225-33234. doi: 10.1073/pnas.2009931117. Epub 2020 Dec 14. Proc Natl Acad Sci U S A. 2020. PMID: 33318190 Free PMC article.
-
Phase separations in oncogenesis, tumor progressions and metastasis: a glance from hallmarks of cancer.J Hematol Oncol. 2023 Dec 18;16(1):123. doi: 10.1186/s13045-023-01522-5. J Hematol Oncol. 2023. PMID: 38110976 Free PMC article. Review.
-
A comparison of four technologies for detecting p53 aggregates in ovarian cancer.Front Oncol. 2022 Sep 8;12:976725. doi: 10.3389/fonc.2022.976725. eCollection 2022. Front Oncol. 2022. PMID: 36158680 Free PMC article.
-
The Etiology and Pathophysiology Genesis of Benign Prostatic Hyperplasia and Prostate Cancer: A New Perspective.Medicines (Basel). 2021 Jun 11;8(6):30. doi: 10.3390/medicines8060030. Medicines (Basel). 2021. PMID: 34208086 Free PMC article. Review.
References
-
- Zhu J., Sammons M. A., Donahue G., Dou Z., Vedadi M., Getlik M., Barsyte-Lovejoy D., Al-awar R., Katona B. W., Shilatifard A., Huang J., Hua X., Arrowsmith C. H., and Berger S. L. (2015) Gain-of-function p53 mutants co-opt chromatin pathways to drive cancer growth. Nature 525, 206–211 10.1038/nature15251 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

