Functional Characterization of microRNA171 Family in Tomato
- PMID: 30621201
- PMCID: PMC6358981
- DOI: 10.3390/plants8010010
Functional Characterization of microRNA171 Family in Tomato
Abstract
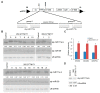
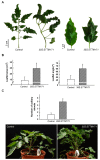
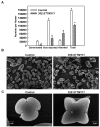
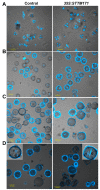
Deeply conserved plant microRNAs (miRNAs) function as pivotal regulators of development. Nevertheless, in the model crop Solanum lycopersicum (tomato) several conserved miRNAs are still poorly annotated and knowledge about their functions is lacking. Here, the tomato miR171 family was functionally analyzed. We found that the tomato genome contains at least 11 SlMIR171 genes that are differentially expressed along tomato development. Downregulation of sly-miR171 in tomato was successfully achieved by transgenic expression of a short tandem target mimic construct (STTM171). Consequently, sly-miR171-targeted mRNAs were upregulated in the silenced plants. Target upregulation was associated with irregular compound leaf development and an increase in the number of axillary branches. A prominent phenotype of STTM171 expressing plants was their male sterility due to a production of a low number of malformed and nonviable pollen. We showed that sly-miR171 was expressed in anthers along microsporogenesis and significantly silenced upon STTM171 expression. Sly-miR171-silenced anthers showed delayed tapetum ontogenesis and reduced callose deposition around the tetrads, both of which together or separately can impair pollen development. Collectively, our results show that sly-miR171 is involved in the regulation of anther development as well as shoot branching and compound leaf morphogenesis.
Keywords: STTM; callose; miR171; pollen; tapetum; tomato.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
LinkOut - more resources
Full Text Sources
Other Literature Sources

