Compositional reorganization of the nucleolus in budding yeast mitosis
- PMID: 30625028
- PMCID: PMC6589692
- DOI: 10.1091/mbc.E18-08-0524
Compositional reorganization of the nucleolus in budding yeast mitosis
Abstract
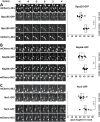
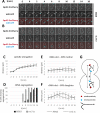
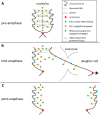
The nucleolus is a membraneless organelle of the nucleus and the site of rRNA synthesis, maturation, and assembly into preribosomal particles. The nucleolus, organized around arrays of rRNA genes (rDNA), dissolves during prophase of mitosis in metazoans, when rDNA transcription ceases, and reforms in telophase, when rDNA transcription resumes. No such dissolution and reformation cycle exists in budding yeast, and the precise course of nucleolar segregation remains unclear. By quantitative live-cell imaging, we observed that the yeast nucleolus is reorganized in its protein composition during mitosis. Daughter cells received equal shares of preinitiation factors, which bind the RNA polymerase I promoter and the rDNA binding barrier protein Fob1, but only about one-third of RNA polymerase I and the processing factors Nop56 and Nsr1. The distribution bias was diminished in nonpolar chromosome segregation events observable in dyn1 mutants. Unequal distribution, however, was enhanced by defects in RNA polymerase I, suggesting that rDNA transcription supports nucleolar segregation. Indeed, quantification of pre-rRNA levels indicated ongoing rDNA transcription in yeast mitosis. These data, together with photobleaching experiments to measure nucleolar protein dynamics in anaphase, consolidate a model that explains the differential partitioning of nucleolar components in budding yeast mitosis.
Figures







Similar articles
-
Targeting of Hmo1 to subcompartments of the budding yeast nucleolus.Mol Biol Cell. 2023 Mar 1;34(3):ar22. doi: 10.1091/mbc.E22-07-0261. Epub 2023 Jan 25. Mol Biol Cell. 2023. PMID: 36696177 Free PMC article.
-
A nucleolus-localized activator of Cdc14 phosphatase supports rDNA segregation in yeast mitosis.Curr Biol. 2008 Jul 8;18(13):1001-5. doi: 10.1016/j.cub.2008.06.025. Curr Biol. 2008. PMID: 18595708
-
Focal accumulation of preribosomes outside the nucleolus during metaphase-anaphase in budding yeast.RNA. 2017 Sep;23(9):1432-1443. doi: 10.1261/rna.061259.117. Epub 2017 Jun 6. RNA. 2017. PMID: 28588079 Free PMC article.
-
A case of selfish nucleolar segregation.Cell Cycle. 2005 Jan;4(1):113-7. doi: 10.4161/cc.4.1.1488. Epub 2005 Jan 21. Cell Cycle. 2005. PMID: 15655374 Review.
-
Cell and molecular biology of nucleolar assembly and disassembly.Int Rev Cytol. 2004;239:99-178. doi: 10.1016/S0074-7696(04)39003-0. Int Rev Cytol. 2004. PMID: 15464853 Review.
Cited by
-
Targeting of Hmo1 to subcompartments of the budding yeast nucleolus.Mol Biol Cell. 2023 Mar 1;34(3):ar22. doi: 10.1091/mbc.E22-07-0261. Epub 2023 Jan 25. Mol Biol Cell. 2023. PMID: 36696177 Free PMC article.
-
Decoding the Nucleolar Role in Meiotic Recombination and Cell Cycle Control: Insights into Cdc14 Function.Int J Mol Sci. 2024 Nov 29;25(23):12861. doi: 10.3390/ijms252312861. Int J Mol Sci. 2024. PMID: 39684572 Free PMC article. Review.
-
Gametogenesis: Exploring an Endogenous Rejuvenation Program to Understand Cellular Aging and Quality Control.Annu Rev Genet. 2022 Nov 30;56:89-112. doi: 10.1146/annurev-genet-080320-025104. Epub 2022 Jul 25. Annu Rev Genet. 2022. PMID: 35878627 Free PMC article. Review.
-
Biomolecular condensates in plant immunity.Cell Host Microbe. 2025 Aug 13;33(8):1276-1290. doi: 10.1016/j.chom.2025.06.014. Cell Host Microbe. 2025. PMID: 40812178 Review.
-
The challenge of staying in shape: nuclear size matters.Curr Genet. 2021 Aug;67(4):605-612. doi: 10.1007/s00294-021-01176-1. Epub 2021 Mar 29. Curr Genet. 2021. PMID: 33779777 Review.
References
-
- Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K. (eds.) (2005). Current Protocols in Molecular Biology, New York: John Wiley & Sons.
-
- Barandun J, Chaker-Margot M, Hunziker M, Molloy KR, Chait BT, Klinge S. (2017). The complete structure of the small-subunit processome. Nat Struct Mol Biol , 944–953. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

