Foxo3a-dependent miR-633 regulates chemotherapeutic sensitivity in gastric cancer by targeting Fas-associated death domain
- PMID: 30628514
- PMCID: PMC6380283
- DOI: 10.1080/15476286.2019.1565665
Foxo3a-dependent miR-633 regulates chemotherapeutic sensitivity in gastric cancer by targeting Fas-associated death domain
Erratum in
-
Correction.RNA Biol. 2019 Aug;16(8):1074. doi: 10.1080/15476286.2019.1609768. Epub 2019 Apr 30. RNA Biol. 2019. PMID: 31035866 Free PMC article. No abstract available.
Abstract
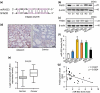
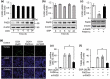
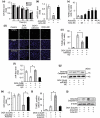
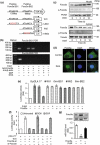
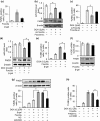
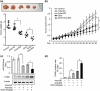
The development of chemotherapeutic drugs resistance such as doxorubicin (DOX) and cisplatin (DDP) is the major barrier in gastric cancer therapy. Emerging evidences reveal that microRNAs (miRNAs) contribute to chemosensitivity. In this study, we investigated the role of miR-633, an oncogenic miRNA, in gastric cancer chemoresistance. In gastric cancer tissue and cell lines, miR-633 expression was highly increased and correlated with down regulation of Fas-associated protein with death domain (FADD). Inhibition of miR-633 significantly increased FADD protein level and enhanced DOX/DDP induced apoptosis in vitro. MiR-633 antagomir administration remarkably decreased tumor growth in combination with DOX in vivo, suggesting that miR-633 targets FADD to block gastric cancer cell death. We found that the promoter region of miR-633 contained putative binding sites for forkhead box O 3 (Foxo3a), which can directly repress miR-633 transcription. In addition, we observed that DOX-induced nuclear accumulation of Foxo3a leaded to the suppression of miR-633 transcription. Together, our study revealed that miR-633/FADD axis played a significant role in the chemoresistance and Foxo3a regulated this pathway in gastric cancer. Thus, miR-633 antagomir resensitized gastric cancer cells to chemotherapy drug and had potentially therapeutic implication.
Keywords: FADD; Foxo3a; chemosensitivity; gastric cancer; miR-633.
Figures

Similar articles
-
Targeted Regulation of FoxO3a by miR-372 to Mediate Gastric Carcinoma Cell Apoptosis and DDP Drug Resistance.Cancer Biother Radiopharm. 2020 Dec;35(10):753-759. doi: 10.1089/cbr.2019.3299. Epub 2020 May 5. Cancer Biother Radiopharm. 2020. Retraction in: Cancer Biother Radiopharm. 2021 Oct;36(8):708. doi: 10.1089/cbr.2019.3299.retract. PMID: 32379491 Retracted.
-
Long Noncoding RNA H19/miR-675 Axis Promotes Gastric Cancer via FADD/Caspase 8/Caspase 3 Signaling Pathway.Cell Physiol Biochem. 2017;42(6):2364-2376. doi: 10.1159/000480028. Epub 2017 Aug 18. Cell Physiol Biochem. 2017. PMID: 28848149
-
miR-20a induces cisplatin resistance of a human gastric cancer cell line via targeting CYLD.Mol Med Rep. 2016 Aug;14(2):1742-50. doi: 10.3892/mmr.2016.5413. Epub 2016 Jun 21. Mol Med Rep. 2016. PMID: 27357419
-
Down-regulated MicroRNAs in Gastric Carcinoma May Be Targets for Therapeutic Intervention and Replacement Therapy.Anticancer Res. 2021 Sep;41(9):4185-4202. doi: 10.21873/anticanres.15223. Epub 2021 Sep 1. Anticancer Res. 2021. PMID: 34475038 Review.
-
MicroRNAs: promising chemoresistance biomarkers in gastric cancer with diagnostic and therapeutic potential.World J Gastroenterol. 2014 Oct 14;20(38):13658-66. doi: 10.3748/wjg.v20.i38.13658. World J Gastroenterol. 2014. PMID: 25320504 Free PMC article. Review.
Cited by
-
The Role of MiRNA in Cancer: Pathogenesis, Diagnosis, and Treatment.Methods Mol Biol. 2022;2257:375-422. doi: 10.1007/978-1-0716-1170-8_18. Methods Mol Biol. 2022. PMID: 34432288
-
Noncoding RNAs in gastric cancer: implications for drug resistance.Mol Cancer. 2020 Mar 19;19(1):62. doi: 10.1186/s12943-020-01185-7. Mol Cancer. 2020. PMID: 32192494 Free PMC article. Review.
-
The FOXO family of transcription factors: key molecular players in gastric cancer.J Mol Med (Berl). 2022 Jul;100(7):997-1015. doi: 10.1007/s00109-022-02219-x. Epub 2022 Jun 10. J Mol Med (Berl). 2022. PMID: 35680690 Review.
-
SIRT1 inhibits chemoresistance and cancer stemness of gastric cancer by initiating an AMPK/FOXO3 positive feedback loop.Cell Death Dis. 2020 Feb 12;11(2):115. doi: 10.1038/s41419-020-2308-4. Cell Death Dis. 2020. PMID: 32051395 Free PMC article.
-
The related miRNAs involved in doxorubicin resistance or sensitivity of various cancers: an update.Cancer Chemother Pharmacol. 2021 Nov;88(5):771-793. doi: 10.1007/s00280-021-04337-8. Epub 2021 Sep 12. Cancer Chemother Pharmacol. 2021. PMID: 34510251 Review.
References
-
- Van Cutsem E, Sagaert X, Topal B, et al. Gastric cancer. Lancet. 2016. November 26;388(10060):2654–2664. PubMed PMID: 27156933; eng. - PubMed
-
- Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016. Mar-Apr;66(2):115–132. PubMed PMID: 26808342; eng. - PubMed
-
- Allemani C, Matsuda T, Di Carlo V, et al. Global surveillance of trends in cancer survival 2000-14 (CONCORD-3): analysis of individual records for 37 513 025 patients diagnosed with one of 18 cancers from 322 population-based registries in 71 countries. Lancet. 2018. March 17;391(10125):1023–1075. PubMed PMID: 29395269; PubMed Central PMCID: PMCPmc5879496. eng. - PMC - PubMed
-
- Tewey KM, Rowe TC, Yang L, et al. Adriamycin-induced DNA damage mediated by mammalian DNA topoisomerase II. Science (New York, NY). 1984. October 26;226(4673):466–468. PubMed PMID: 6093249; eng. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous
