Secreted amyloid-β precursor protein functions as a GABABR1a ligand to modulate synaptic transmission
- PMID: 30630900
- PMCID: PMC6366617
- DOI: 10.1126/science.aao4827
Secreted amyloid-β precursor protein functions as a GABABR1a ligand to modulate synaptic transmission
Abstract
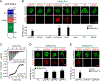
Amyloid-β precursor protein (APP) is central to the pathogenesis of Alzheimer's disease, yet its physiological function remains unresolved. Accumulating evidence suggests that APP has a synaptic function mediated by an unidentified receptor for secreted APP (sAPP). Here we show that the sAPP extension domain directly bound the sushi 1 domain specific to the γ-aminobutyric acid type B receptor subunit 1a (GABABR1a). sAPP-GABABR1a binding suppressed synaptic transmission and enhanced short-term facilitation in mouse hippocampal synapses via inhibition of synaptic vesicle release. A 17-amino acid peptide corresponding to the GABABR1a binding region within APP suppressed in vivo spontaneous neuronal activity in the hippocampus of anesthetized Thy1-GCaMP6s mice. Our findings identify GABABR1a as a synaptic receptor for sAPP and reveal a physiological role for sAPP in regulating GABABR1a function to modulate synaptic transmission.
Copyright © 2019 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Conflict of interest statement
Figures


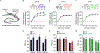


Comment in
-
Neuronal function of Alzheimer's protein.Science. 2019 Jan 11;363(6423):123-124. doi: 10.1126/science.aaw0636. Science. 2019. PMID: 30630916 No abstract available.
-
Revealing a receptor for secreted APP.Nat Rev Neurosci. 2019 Mar;20(3):129. doi: 10.1038/s41583-019-0136-2. Nat Rev Neurosci. 2019. PMID: 30733610 No abstract available.
-
Secreted APP Modulates Synaptic Activity: A Novel Target for Therapeutic Intervention?Neuron. 2019 Feb 20;101(4):557-559. doi: 10.1016/j.neuron.2019.01.058. Neuron. 2019. PMID: 30790537
Similar articles
-
Molecular mechanism of secreted amyloid-β precursor protein in binding and modulating GABABR1a.Chem Sci. 2021 Mar 16;12(17):6107-6116. doi: 10.1039/d0sc06946a. Chem Sci. 2021. PMID: 33996007 Free PMC article.
-
Amyloid Precursor Protein (APP) and GABAergic Neurotransmission.Cells. 2019 Jun 6;8(6):550. doi: 10.3390/cells8060550. Cells. 2019. PMID: 31174368 Free PMC article.
-
Increased activity-regulating and neuroprotective efficacy of alpha-secretase-derived secreted amyloid precursor protein conferred by a C-terminal heparin-binding domain.J Neurochem. 1996 Nov;67(5):1882-96. doi: 10.1046/j.1471-4159.1996.67051882.x. J Neurochem. 1996. PMID: 8863493
-
The role of APP and APLP for synaptic transmission, plasticity, and network function: lessons from genetic mouse models.Exp Brain Res. 2012 Apr;217(3-4):435-40. doi: 10.1007/s00221-011-2894-6. Epub 2011 Oct 18. Exp Brain Res. 2012. PMID: 22006270 Review.
-
Perturbed endoplasmic reticulum function, synaptic apoptosis and the pathogenesis of Alzheimer's disease.Biochem Soc Symp. 2001;(67):151-62. doi: 10.1042/bss0670151. Biochem Soc Symp. 2001. PMID: 11447832 Review.
Cited by
-
GABAergic Inhibitory Interneuron Deficits in Alzheimer's Disease: Implications for Treatment.Front Neurosci. 2020 Jun 30;14:660. doi: 10.3389/fnins.2020.00660. eCollection 2020. Front Neurosci. 2020. PMID: 32714136 Free PMC article. Review.
-
The Impairment of Blood-Brain Barrier in Alzheimer's Disease: Challenges and Opportunities with Stem Cells.Int J Mol Sci. 2022 Sep 4;23(17):10136. doi: 10.3390/ijms231710136. Int J Mol Sci. 2022. PMID: 36077533 Free PMC article. Review.
-
Symptomatic, Genetic, and Mechanistic Overlaps between Autism and Alzheimer's Disease.Biomolecules. 2021 Nov 4;11(11):1635. doi: 10.3390/biom11111635. Biomolecules. 2021. PMID: 34827633 Free PMC article. Review.
-
Does HIV infection contribute to increased beta-amyloid synthesis and plaque formation leading to neurodegeneration and Alzheimer's disease?J Neurovirol. 2019 Oct;25(5):634-647. doi: 10.1007/s13365-019-00732-3. Epub 2019 Mar 13. J Neurovirol. 2019. PMID: 30868421 Review.
-
Structure of the photoreceptor synaptic assembly of the extracellular matrix protein pikachurin with the orphan receptor GPR179.Sci Signal. 2023 Jul 25;16(795):eadd9539. doi: 10.1126/scisignal.add9539. Epub 2023 Jul 25. Sci Signal. 2023. PMID: 37490546 Free PMC article.
References
-
- Goldgaber D, Lerman M, McBride OW, Saffiotti U, Gajdusek DC, Characterization and Chromosomal Localization of a cDNA Encoding Brain Amyloid of Alzheimer’s Disease. Science. 235, 877–880 (1987). - PubMed
-
- N. R. Tanzi RE, Gusella JF, Watkins PC, Bruns GA, St George-Hyslop P, Van Keuren ML, Patterson D, Pagan S, Kurnit DM, Amyloid beta protein gene: cDNA, mRNA distribution, and genetic linkage near the Alzheimer locus. Science. 235 (1987). - PubMed
-
- Kang J et al., The precursor of Alzheimer’s disease amyloid A4 protein resembles a cell-surface receptor. Nature. 325 (1987), pp. 733–6. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

