Expression of novel long noncoding RNAs defines virus-specific effector and memory CD8+ T cells
- PMID: 30643116
- PMCID: PMC6331603
- DOI: 10.1038/s41467-018-07956-7
Expression of novel long noncoding RNAs defines virus-specific effector and memory CD8+ T cells
Abstract
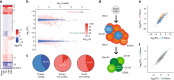
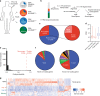
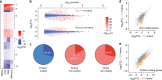
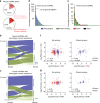
In response to viral infection, CD8+ T cells undergo expansion and differentiate into distinct classes of effector cells. After clearance of the virus, a small population of long-lived memory cells persists. Comprehensive studies have defined the protein-coding transcriptional changes associated with this process. Here we expand on this prior work by performing RNA-sequencing to identify changes in long noncoding RNA (lncRNA) expression in human and mouse CD8+ T cells responding to viral infection. We identify hundreds of unannotated lncRNAs and show that expression profiles of both known and novel lncRNAs are sufficient to define naive, effector, and memory CD8+ T cell subsets, implying that they may be involved in fate decisions during antigen-driven differentiation. Additionally, in comparing mouse and human lncRNA expression, we find that lncRNAs with conserved sequence undergo similar changes in expression in the two species, suggesting an evolutionarily conserved role for lncRNAs during CD8+ T cell differentiation.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Transcriptional repressor ZEB2 promotes terminal differentiation of CD8+ effector and memory T cell populations during infection.J Exp Med. 2015 Nov 16;212(12):2027-39. doi: 10.1084/jem.20150194. Epub 2015 Oct 26. J Exp Med. 2015. PMID: 26503445 Free PMC article.
-
The long noncoding RNA Morrbid regulates CD8 T cells in response to viral infection.Proc Natl Acad Sci U S A. 2019 Jun 11;116(24):11916-11925. doi: 10.1073/pnas.1819457116. Epub 2019 May 28. Proc Natl Acad Sci U S A. 2019. PMID: 31138702 Free PMC article.
-
Differentiation of CD8 memory T cells depends on Foxo1.J Exp Med. 2013 Jun 3;210(6):1189-200. doi: 10.1084/jem.20130392. Epub 2013 May 27. J Exp Med. 2013. PMID: 23712431 Free PMC article.
-
Non-coding RNAs in CD8 T cell biology.Mol Immunol. 2020 Apr;120:67-73. doi: 10.1016/j.molimm.2020.01.023. Epub 2020 Feb 18. Mol Immunol. 2020. PMID: 32085976 Free PMC article. Review.
-
CD8 T cell dysfunction during chronic viral infection.Curr Opin Immunol. 2007 Aug;19(4):408-15. doi: 10.1016/j.coi.2007.06.004. Epub 2007 Jul 25. Curr Opin Immunol. 2007. PMID: 17656078 Review.
Cited by
-
RDUR, a lncRNA, Promotes Innate Antiviral Responses and Provides Feedback Control of NF-κB Activation.Front Immunol. 2021 May 14;12:672165. doi: 10.3389/fimmu.2021.672165. eCollection 2021. Front Immunol. 2021. PMID: 34054851 Free PMC article.
-
Downregulation of the Long Noncoding RNA IALNCR Targeting MAPK8/JNK1 Promotes Apoptosis and Antagonizes Bovine Viral Diarrhea Virus Replication in Host Cells.J Virol. 2022 Sep 14;96(17):e0111322. doi: 10.1128/jvi.01113-22. Epub 2022 Aug 22. J Virol. 2022. PMID: 35993735 Free PMC article.
-
Transcriptomics of chicken cecal tonsils and intestine after infection with low pathogenic avian influenza virus H9N2.Sci Rep. 2021 Oct 14;11(1):20462. doi: 10.1038/s41598-021-99182-3. Sci Rep. 2021. PMID: 34650121 Free PMC article.
-
Transcriptome Analysis Identifies SenZfp536, a Sense LncRNA that Suppresses Self-renewal of Cortical Neural Progenitors.Neurosci Bull. 2021 Feb;37(2):183-200. doi: 10.1007/s12264-020-00607-2. Epub 2020 Nov 16. Neurosci Bull. 2021. PMID: 33196962 Free PMC article.
-
Prospective Study of Long Noncoding RNA, MGAT3-AS1, and Viremia of BK Polyomavirus and Cytomegalovirus in Living Donor Renal Transplant Recipients.Kidney Int Rep. 2020 Sep 6;5(12):2218-2227. doi: 10.1016/j.ekir.2020.09.005. eCollection 2020 Dec. Kidney Int Rep. 2020. PMID: 33305115 Free PMC article.
References
-
- Kaech SM, Hemby S, Kersh E, Ahmed R. Molecular and functional profiling of memory CD8 T. Cell Differ. Cell. 2002;111:837–851. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

