Segmented poly(A) tails significantly reduce recombination of plasmid DNA without affecting mRNA translation efficiency or half-life
- PMID: 30647100
- PMCID: PMC6426288
- DOI: 10.1261/rna.069286.118
Segmented poly(A) tails significantly reduce recombination of plasmid DNA without affecting mRNA translation efficiency or half-life
Abstract
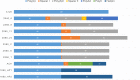
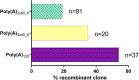
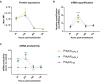
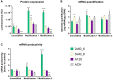
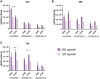
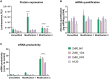
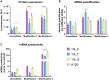
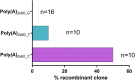
Extensive research in the past decade has brought mRNA closer to the clinical realization of its therapeutic potential. One common structural feature for all cellular messenger RNAs is a poly(A) tail, which can either be brought in cotranscriptionally via the DNA template (plasmid- or PCR-based) or added to the mRNA in a post-transcriptional enzymatic process. Plasmids containing poly(A) regions recombine in E. coli, resulting in extensive shortening of the poly(A) tail. Using a segmented poly(A) approach, we could significantly reduce recombination of plasmids in E. coli without any negative effect on mRNA half-life and protein expression. This effect was independent of the coding sequence. A segmented poly(A) tail is characterized in that it consists of at least two A-containing elements, each defined as a nucleotide sequence consisting of 40-60 adenosines, separated by a spacer element of different length. Furthermore, reducing the spacer length between the poly(A) segments resulted in higher translation efficiencies compared to homogeneous poly(A) tail and reduced recombination (depending upon the choice of spacer nucleotide). Our results demonstrate the superior potential of segmented poly(A) tails compared to the conventionally used homogeneous poly(A) tails with respect to recombination of the plasmids and the resulting mRNA performance (half-life and translational efficiency).
Keywords: mRNA therapeutics; plasmid recombination; poly(A) tail; transcript therapy.
© 2019 Trepotec et al.; Published by Cold Spring Harbor Laboratory Press for the RNA Society.
Figures

Similar articles
-
Decoupled degradation and translation enables noise modulation by poly(A) tails.Cell Syst. 2024 Jun 19;15(6):526-543.e7. doi: 10.1016/j.cels.2024.05.004. Cell Syst. 2024. PMID: 38901403
-
Messenger RNA stability in Saccharomyces cerevisiae: the influence of translation and poly(A) tail length.Nucleic Acids Res. 1987 Mar 25;15(6):2417-29. doi: 10.1093/nar/15.6.2417. Nucleic Acids Res. 1987. PMID: 3550698 Free PMC article.
-
Poly A tail length analysis of in vitro transcribed mRNA by LC-MS.Anal Bioanal Chem. 2018 Feb;410(6):1667-1677. doi: 10.1007/s00216-017-0840-6. Epub 2018 Jan 8. Anal Bioanal Chem. 2018. PMID: 29313076
-
Tales of Detailed Poly(A) Tails.Trends Cell Biol. 2019 Mar;29(3):191-200. doi: 10.1016/j.tcb.2018.11.002. Epub 2018 Nov 29. Trends Cell Biol. 2019. PMID: 30503240 Free PMC article. Review.
-
Why, when and how does the poly(A) tail shorten during mRNA translation?Int J Biochem. 1993 Mar;25(3):287-95. doi: 10.1016/0020-711x(93)90615-l. Int J Biochem. 1993. PMID: 8462718 Review.
Cited by
-
mRNA Therapeutic Modalities Design, Formulation and Manufacturing under Pharma 4.0 Principles.Biomedicines. 2021 Dec 27;10(1):50. doi: 10.3390/biomedicines10010050. Biomedicines. 2021. PMID: 35052730 Free PMC article. Review.
-
mRNA-Based Vaccines and Mode of Action.Curr Top Microbiol Immunol. 2022;440:1-30. doi: 10.1007/82_2020_230. Curr Top Microbiol Immunol. 2022. PMID: 33591423 Review.
-
Supercoiled DNA percentage: A key in-process control of linear DNA template for mRNA drug substance manufacturing.Mol Ther Nucleic Acids. 2024 May 20;35(2):102223. doi: 10.1016/j.omtn.2024.102223. eCollection 2024 Jun 11. Mol Ther Nucleic Acids. 2024. PMID: 38948330 Free PMC article.
-
A (RP)UHPLC/UV analytical method to quantify dsRNA during the mRNA vaccine manufacturing process.Anal Methods. 2024 Aug 1;16(30):5146-5153. doi: 10.1039/d4ay00560k. Anal Methods. 2024. PMID: 39011770 Free PMC article.
-
Liquid Chromatography-Mass Spectrometry-Based Qualitative Profiling of mRNA Therapeutic Reagents Using Stable Isotope-Labeled Standards Followed by the Automatic Quantitation Software Ariadne.Anal Chem. 2023 Jan 17;95(2):1366-1375. doi: 10.1021/acs.analchem.2c04323. Epub 2022 Dec 27. Anal Chem. 2023. PMID: 36574727 Free PMC article.
References
-
- Bahl K, Senn JJ, Yuzhakov O, Bulychev A, Brito LA, Hassett KJ, Laska ME, Smith M, Almarsson Ö, Thompson J, et al. 2017. Preclinical and clinical demonstration of immunogenicity by mRNA vaccines against H10N8 and H7N9 influenza viruses. Mol Ther 25: 1316–1327. 10.1016/j.ymthe.2017.03.035 - DOI - PMC - PubMed
-
- Balmayor ER, Geiger JP, Aneja MK, Berezhanskyy T, Utzinger M, Mykhaylyk O, Rudolph C, Plank C. 2016. Chemically modified RNA induces osteogenesis of stem cells and human tissue explants as well as accelerates bone healing in rats. Biomaterials 87: 131–146. 10.1016/j.biomaterials.2016.02.018 - DOI - PubMed
-
- Balmayor ER, Geiger JP, Koch C, Aneja MK, van Griensven M, Rudolph C, Plank C. 2017. Modified mRNA for BMP-2 in combination with biomaterials serves as a transcript-activated matrix for effectively inducing osteogenic pathways in stem cells. Stem Cells Dev 26: 25–34. 10.1089/scd.2016.0171 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
