Internal Gain Modulations, But Not Changes in Stimulus Contrast, Preserve the Neural Code
- PMID: 30647148
- PMCID: PMC6391566
- DOI: 10.1523/JNEUROSCI.2012-18.2019
Internal Gain Modulations, But Not Changes in Stimulus Contrast, Preserve the Neural Code
Abstract
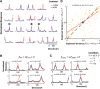
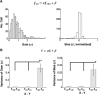
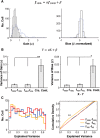
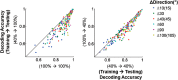
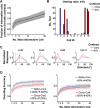
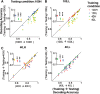
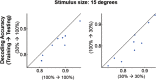
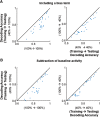
Neurons in primary visual cortex are strongly modulated both by stimulus contrast and by fluctuations of internal inputs. An important question is whether the population code is preserved under these conditions. Changes in stimulus contrast are thought to leave the population code invariant, whereas the effect of internal gain modulations remains unknown. To address these questions we studied how the direction-of-motion of oriented gratings is encoded in layer 2/3 primary visual cortex of mouse (with C57BL/6 background, of either sex). We found that, because contrast gain responses across cells are heterogeneous, a change in contrast alters the information distribution profile across cells leading to a violation of contrast invariance. Remarkably, internal input fluctuations that cause commensurate firing rate modulations at the single-cell level result in more homogeneous gain responses, respecting population code invariance. These observations argue that the brain strives to maintain the stability of the neural code in the face of fluctuating internal inputs.SIGNIFICANCE STATEMENT Neuronal responses are modulated both by stimulus contrast and by the spontaneous fluctuation of internal inputs. It is not well understood how these different types of input impact the population code. Specifically, it is important to understand whether the neural code stays invariant in the face of significant internal input modulations. Here, we show that changes in stimulus contrast lead to different optimal population codes, whereas spontaneous internal input fluctuations leave the population code invariant. This is because spontaneous internal input fluctuations modulate the gain of neuronal responses more homogeneously across cells compared to changes in stimulus contrast.
Keywords: brain states; mouse visual cortex; population codes; two-photon calcium imaging; visual contrast.
Copyright © 2019 the authors 0270-6474/19/391671-17$15.00/0.
Figures






References
-
- Bishop CM. (2006) Pattern recognition and machine learning. New York: Springer.
-
- Boyd SP, Vandenberghe L (2004) Convex optimization. Cambridge, UK; New York: Cambridge UP.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
