N-Cadherin-Expressing Bone and Marrow Stromal Progenitor Cells Maintain Reserve Hematopoietic Stem Cells
- PMID: 30650358
- PMCID: PMC6890378
- DOI: 10.1016/j.celrep.2018.12.093
N-Cadherin-Expressing Bone and Marrow Stromal Progenitor Cells Maintain Reserve Hematopoietic Stem Cells
Abstract
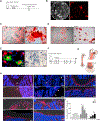
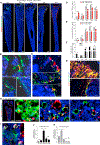
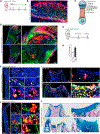
Regulation of hematopoietic stem cells (HSCs) by bone marrow (BM) niches has been extensively studied; however, whether and how HSC subpopulations are distinctively regulated by BM niches remain unclear. Here, we functionally distinguished reserve HSCs (rHSCs) from primed HSCs (pHSCs) based on their response to chemotherapy and examined how they are dichotomously regulated by BM niches. Both pHSCs and rHSCs supported long-term hematopoiesis in homeostasis; however, pHSCs were sensitive but rHSCs were resistant to chemotherapy. Surviving rHSCs restored the HSC pool and supported hematopoietic regeneration after chemotherapy. The rHSCs were preferentially maintained in the endosteal region that enriches N-cadherin+ (N-cad+) bone-lining cells in homeostasis and post-chemotherapy. N-cad+ cells were functional bone and marrow stromal progenitor cells (BMSPCs), giving rise to osteoblasts, adipocytes, and chondrocytes in vitro and in vivo. Finally, ablation of N-cad+ niche cells or deletion of SCF from N-cad+ niche cells impaired rHSC maintenance during homeostasis and regeneration.
Keywords: MSC; N-cadherin; endosteum; niche; reserve stem cell; stress response.
Copyright © 2018 Stowers Institute for Medical Research. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
DECLARATION OF INTERESTS
The authors declare no competing interests.
Figures




Comment in
-
Dynamic Changes in the Niche with N-Cadherin Revisited: The HSC "Niche Herein".Cell Stem Cell. 2019 Mar 7;24(3):355-356. doi: 10.1016/j.stem.2019.02.007. Cell Stem Cell. 2019. PMID: 30849365
References
-
- Arai F, Hirao A, Ohmura M, Sato H, Matsuoka S, Takubo K, Ito K, Koh GY, and Suda T. (2004). Tie2/angiopoietin-1 signaling regulates hematopoietic stem cell quiescence in the bone marrow niche. Cell 118, 149–161. - PubMed
-
- Arnold MA, Kim Y, Czubryt MP, Phan D, McAnally J, Qi X, Shelton JM, Richardson JA, Bassel-Duby R, and Olson EN (2007). MEF2C transcription factor controls chondrocyte hypertrophy and bone development. Dev. Cell 12, 377–389. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

