Expanding Repertoire of Plant Positive-Strand RNA Virus Proteases
- PMID: 30650571
- PMCID: PMC6357015
- DOI: 10.3390/v11010066
Expanding Repertoire of Plant Positive-Strand RNA Virus Proteases
Abstract
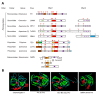
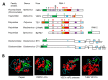
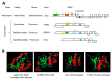
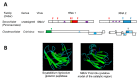
Many plant viruses express their proteins through a polyprotein strategy, requiring the acquisition of protease domains to regulate the release of functional mature proteins and/or intermediate polyproteins. Positive-strand RNA viruses constitute the vast majority of plant viruses and they are diverse in their genomic organization and protein expression strategies. Until recently, proteases encoded by positive-strand RNA viruses were described as belonging to two categories: (1) chymotrypsin-like cysteine and serine proteases and (2) papain-like cysteine protease. However, the functional characterization of plant virus cysteine and serine proteases has highlighted their diversity in terms of biological activities, cleavage site specificities, regulatory mechanisms, and three-dimensional structures. The recent discovery of a plant picorna-like virus glutamic protease with possible structural similarities with fungal and bacterial glutamic proteases also revealed new unexpected sources of protease domains. We discuss the variety of plant positive-strand RNA virus protease domains. We also highlight possible evolution scenarios of these viral proteases, including evidence for the exchange of protease domains amongst unrelated viruses.
Keywords: protease specificity; protease structure; proteolytic processing; viral proteases; virus evolution.
Conflict of interest statement
“The authors declare no conflict of interest.”
Figures
Similar articles
-
Strawberry Mottle Virus (Family Secoviridae, Order Picornavirales) Encodes a Novel Glutamic Protease To Process the RNA2 Polyprotein at Two Cleavage Sites.J Virol. 2019 Feb 19;93(5):e01679-18. doi: 10.1128/JVI.01679-18. Print 2019 Mar 1. J Virol. 2019. PMID: 30541838 Free PMC article.
-
Computer-assisted assignment of functional domains in the nonstructural polyprotein of hepatitis E virus: delineation of an additional group of positive-strand RNA plant and animal viruses.Proc Natl Acad Sci U S A. 1992 Sep 1;89(17):8259-63. doi: 10.1073/pnas.89.17.8259. Proc Natl Acad Sci U S A. 1992. PMID: 1518855 Free PMC article.
-
Putative papain-related thiol proteases of positive-strand RNA viruses. Identification of rubi- and aphthovirus proteases and delineation of a novel conserved domain associated with proteases of rubi-, alpha- and coronaviruses.FEBS Lett. 1991 Aug 19;288(1-2):201-5. doi: 10.1016/0014-5793(91)81034-6. FEBS Lett. 1991. PMID: 1652473 Free PMC article.
-
Cysteine proteases of positive strand RNA viruses and chymotrypsin-like serine proteases. A distinct protein superfamily with a common structural fold.FEBS Lett. 1989 Jan 30;243(2):103-14. doi: 10.1016/0014-5793(89)80109-7. FEBS Lett. 1989. PMID: 2645167 Review.
-
Evolution and taxonomy of positive-strand RNA viruses: implications of comparative analysis of amino acid sequences.Crit Rev Biochem Mol Biol. 1993;28(5):375-430. doi: 10.3109/10409239309078440. Crit Rev Biochem Mol Biol. 1993. PMID: 8269709 Review.
Cited by
-
Re-examination of nepovirus polyprotein cleavage sites highlights the diverse specificities and evolutionary relationships of nepovirus 3C-like proteases.Arch Virol. 2022 Dec;167(12):2529-2543. doi: 10.1007/s00705-022-05564-x. Epub 2022 Aug 30. Arch Virol. 2022. PMID: 36042138 Free PMC article.
-
Potyviral Helper-Component Protease: Multifaced Functions and Interactions with Host Proteins.Plants (Basel). 2024 Apr 29;13(9):1236. doi: 10.3390/plants13091236. Plants (Basel). 2024. PMID: 38732454 Free PMC article. Review.
-
Abscisic Acid Connects Phytohormone Signaling with RNA Metabolic Pathways and Promotes an Antiviral Response that Is Evaded by a Self-Controlled RNA Virus.Plant Commun. 2020 Sep 14;1(5):100099. doi: 10.1016/j.xplc.2020.100099. Epub 2020 Jul 7. Plant Commun. 2020. PMID: 32984814 Free PMC article.
-
CSV2018: The 2nd Symposium of the Canadian Society for Virology.Viruses. 2019 Jan 18;11(1):79. doi: 10.3390/v11010079. Viruses. 2019. PMID: 30669273 Free PMC article.
-
A Novel Rice Curl Dwarf-Associated Picornavirus Encodes a 3C Serine Protease Recognizing Uncommon EPT/S Cleavage Sites.Front Microbiol. 2021 Oct 13;12:757451. doi: 10.3389/fmicb.2021.757451. eCollection 2021. Front Microbiol. 2021. PMID: 34721366 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

