Dystrophin Deficiency Leads to Genomic Instability in Human Pluripotent Stem Cells via NO Synthase-Induced Oxidative Stress
- PMID: 30650618
- PMCID: PMC6356905
- DOI: 10.3390/cells8010053
Dystrophin Deficiency Leads to Genomic Instability in Human Pluripotent Stem Cells via NO Synthase-Induced Oxidative Stress
Abstract
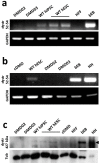
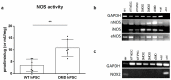
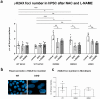
Recent data on Duchenne muscular dystrophy (DMD) show myocyte progenitor's involvement in the disease pathology often leading to the DMD patient's death. The molecular mechanism underlying stem cell impairment in DMD has not been described. We created dystrophin-deficient human pluripotent stem cell (hPSC) lines by reprogramming cells from two DMD patients, and also by introducing dystrophin mutation into human embryonic stem cells via CRISPR/Cas9. While dystrophin is expressed in healthy hPSC, its deficiency in DMD hPSC lines induces the release of reactive oxygen species (ROS) through dysregulated activity of all three isoforms of nitric oxide synthase (further abrev. as, NOS). NOS-induced ROS release leads to DNA damage and genomic instability in DMD hPSC. We were able to reduce both the ROS release as well as DNA damage to the level of wild-type hPSC by inhibiting NOS activity.
Keywords: DMD; NO synthases; ROS; dystrophin; genome stability; pluripotent stem cells.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





Similar articles
-
A Single CRISPR-Cas9 Deletion Strategy that Targets the Majority of DMD Patients Restores Dystrophin Function in hiPSC-Derived Muscle Cells.Cell Stem Cell. 2016 Apr 7;18(4):533-40. doi: 10.1016/j.stem.2016.01.021. Epub 2016 Feb 11. Cell Stem Cell. 2016. PMID: 26877224 Free PMC article.
-
Absence of full-length dystrophin impairs normal maturation and contraction of cardiomyocytes derived from human-induced pluripotent stem cells.Cardiovasc Res. 2020 Feb 1;116(2):368-382. doi: 10.1093/cvr/cvz109. Cardiovasc Res. 2020. PMID: 31049579 Free PMC article.
-
Fibre-related nitric oxide synthase (NOS) in Duchenne muscular dystrophy.Acta Histochem. 2007;109(3):228-36. doi: 10.1016/j.acthis.2007.01.001. Epub 2007 Feb 20. Acta Histochem. 2007. PMID: 17313973
-
The involvement of oxidative stress in determining the severity and progress of pathological processes in dystrophin-deficient muscles.Acta Biochim Pol. 2005;52(2):449-52. Epub 2005 May 25. Acta Biochim Pol. 2005. PMID: 15990924 Review.
-
Therapeutic Applications of CRISPR/Cas for Duchenne Muscular Dystrophy.Curr Gene Ther. 2017;17(4):301-308. doi: 10.2174/1566523217666171121165046. Curr Gene Ther. 2017. PMID: 29173172 Review.
Cited by
-
Decoding the Gene Regulatory Network of Muscle Stem Cells in Mouse Duchenne Muscular Dystrophy: Revelations from Single-Nuclei RNA Sequencing Analysis.Int J Mol Sci. 2023 Aug 5;24(15):12463. doi: 10.3390/ijms241512463. Int J Mol Sci. 2023. PMID: 37569835 Free PMC article.
-
Calcium handling maturation and adaptation to increased substrate stiffness in human iPSC-derived cardiomyocytes: The impact of full-length dystrophin deficiency.Front Physiol. 2022 Nov 7;13:1030920. doi: 10.3389/fphys.2022.1030920. eCollection 2022. Front Physiol. 2022. PMID: 36419836 Free PMC article.
-
Advances in Stem Cell Modeling of Dystrophin-Associated Disease: Implications for the Wider World of Dilated Cardiomyopathy.Front Physiol. 2020 May 12;11:368. doi: 10.3389/fphys.2020.00368. eCollection 2020. Front Physiol. 2020. PMID: 32477154 Free PMC article. Review.
-
Tumor Necrosis Factor Receptor SF10A (TNFRSF10A) SNPs Correlate With Corticosteroid Response in Duchenne Muscular Dystrophy.Front Genet. 2020 Jul 3;11:605. doi: 10.3389/fgene.2020.00605. eCollection 2020. Front Genet. 2020. PMID: 32719714 Free PMC article.
-
Restoration of Dystrophin Expression in Mdx-Derived Muscle Progenitor Cells Using CRISPR/Cas9 System and Homology-Directed Repair Technology.Methods Mol Biol. 2023;2587:455-464. doi: 10.1007/978-1-0716-2772-3_23. Methods Mol Biol. 2023. PMID: 36401043
References
-
- Tidball J.G., Albrecht D.E., Lokensgard B.E., Spencer M.J. Apoptosis precedes necrosis of dystrophin-deficient muscle. (Pt 6)J. Cell Sci. 1995;108:2197–2204. - PubMed
-
- Mikhaĭlov V.M., Komarov S.A., Nilova V.K., Shteĭn G.I., Baranov V.S. Ultrastructural and morphometrical analysis of apoptosis stages in cardiomyocytes of MDX mice. Tsitologiia. 2001;43:729–737. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

