Resistance to oxidative stress by inner membrane protein ElaB is regulated by OxyR and RpoS
- PMID: 30656833
- PMCID: PMC6389858
- DOI: 10.1111/1751-7915.13369
Resistance to oxidative stress by inner membrane protein ElaB is regulated by OxyR and RpoS
Abstract
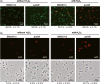
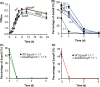
C-tail anchored inner membrane proteins are a family of proteins that contain a C-terminal transmembrane domain but lack an N-terminal signal sequence for membrane targeting. They are widespread in eukaryotes and prokaryotes and play critical roles in membrane traffic, apoptosis and protein translocation in eukaryotes. Recently, we identified and characterized in Escherichia coli a new C-tail anchored inner membrane, ElaB, which is regulated by the stationary phase sigma factor RpoS. ElaB is important for resistance to oxidative stress but the exact mechanism is unclear. Here, we show that ElaB functions as part of the adaptive oxidative stress response by maintaining membrane integrity. Production of ElaB is induced by oxidative stress at the transcriptional level. Moreover, elaB expression is also regulated by the key regulator OxyR via an OxyR binding site in the promoter of elaB. OxyR induces the expression of elaB in the exponential growth phase, while excess OxyR reduces elaB expression in an RpoS-dependent way in the stationary phase. In addition, deletion of elaB reduced fitness compared to wild-type cells after prolonged incubation. Therefore, we determined how ElaB is regulated under oxidative stress: RpoS and OxyR coordinately control the expression of inner membrane protein ElaB.
© 2019 The Authors. Microbial Biotechnology published by John Wiley & Sons Ltd and Society for Applied Microbiology.
Conflict of interest statement
The authors declare no competing financial interests.
Figures



References
-
- Altuvia, S. , Almiron, M. , Huisman, G. , Kolter, R. , and Storz, G. (1994) The dps promoter is activated by OxyR during growth and by IHF and a σS in stationary phase. Mol Microbiol 13: 265–272. - PubMed
-
- Becker‐Hapak, M. , and Eisenstark, A. (1995) Role of rpoS in the regulation of glutathione oxidoreductase (gor) in Escherichia coli . FEMS Microbiol Lett 134: 39–44. - PubMed
-
- Bienert, G.P. , Schjoerring, J.K. , and Jahn, T.P. (2006) Membrane transport of hydrogen peroxide. Biochim Biophys Acta 1758: 994–1003. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

