Sucrose Metabolism in Haloarchaea: Reassessment Using Genomics, Proteomics, and Metagenomics
- PMID: 30658981
- PMCID: PMC6414365
- DOI: 10.1128/AEM.02935-18
Sucrose Metabolism in Haloarchaea: Reassessment Using Genomics, Proteomics, and Metagenomics
Abstract
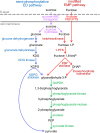
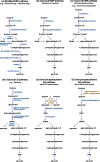
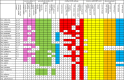
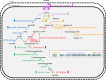
The canonical pathway for sucrose metabolism in haloarchaea utilizes a modified Embden-Meyerhof-Parnas pathway (EMP), in which ketohexokinase and 1-phosphofructokinase phosphorylate fructose released from sucrose hydrolysis. However, our survey of haloarchaeal genomes determined that ketohexokinase and 1-phosphofructokinase genes were not present in all species known to utilize fructose and sucrose, thereby indicating that alternative mechanisms exist for fructose metabolism. A fructokinase gene was identified in the majority of fructose- and sucrose-utilizing species, whereas only a small number possessed a ketohexokinase gene. Analysis of a range of hypersaline metagenomes revealed that haloarchaeal fructokinase genes were far more abundant (37 times) than haloarchaeal ketohexokinase genes. We used proteomic analysis of Halohasta litchfieldiae (which encodes fructokinase) and identified changes in protein abundance that relate to growth on sucrose. Proteins inferred to be involved in sucrose metabolism included fructokinase, a carbohydrate primary transporter, a putative sucrose hydrolase, and two uncharacterized carbohydrate-related proteins encoded in the same gene cluster as fructokinase and the transporter. Homologs of these proteins were present in the genomes of all haloarchaea that use sugars for growth. Enzymes involved in the semiphosphorylative Entner-Doudoroff pathway also had higher abundances in sucrose-grown H. litchfieldiae cells, consistent with this pathway functioning in the catabolism of the glucose moiety of sucrose. The study revises the current understanding of fundamental pathways for sugar utilization in haloarchaea and proposes alternatives to the modified EMP pathway used by haloarchaea for sucrose and fructose utilization.IMPORTANCE Our ability to infer the function that microorganisms perform in the environment is predicated on assumptions about metabolic capacity. When genomic or metagenomic data are used, metabolic capacity is inferred from genetic potential. Here, we investigate the pathways by which haloarchaea utilize sucrose. The canonical haloarchaeal pathway for fructose metabolism involving ketohexokinase occurs only in a small proportion of haloarchaeal genomes and is underrepresented in metagenomes. Instead, fructokinase genes are present in the majority of genomes/metagenomes. In addition to genomic and metagenomic analyses, we used proteomic analysis of Halohasta litchfieldiae (which encodes fructokinase but lacks ketohexokinase) and identified changes in protein abundance that related to growth on sucrose. In this way, we identified novel proteins implicated in sucrose metabolism in haloarchaea, comprising a transporter and various catabolic enzymes (including proteins that are annotated as hypothetical).
Keywords: archaea.
Copyright © 2019 American Society for Microbiology.
Figures

Similar articles
-
Fructose metabolism in Chromohalobacter salexigens: interplay between the Embden-Meyerhof-Parnas and Entner-Doudoroff pathways.Microb Cell Fact. 2019 Aug 13;18(1):134. doi: 10.1186/s12934-019-1178-x. Microb Cell Fact. 2019. PMID: 31409414 Free PMC article.
-
Identification and characterization of a novel type of ketohexokinase from the haloarchaeon Haloferax volcanii.FEMS Microbiol Lett. 2024 Jan 9;371:fnae026. doi: 10.1093/femsle/fnae026. FEMS Microbiol Lett. 2024. PMID: 38587824
-
Sugar alcohol degradation in Archaea: uptake and degradation of mannitol and sorbitol in Haloarcula hispanica.Extremophiles. 2024 Oct 28;28(3):48. doi: 10.1007/s00792-024-01365-z. Extremophiles. 2024. PMID: 39466404 Free PMC article.
-
Unusual pathways and enzymes of central carbohydrate metabolism in Archaea.Curr Opin Microbiol. 2005 Dec;8(6):695-705. doi: 10.1016/j.mib.2005.10.014. Epub 2005 Oct 26. Curr Opin Microbiol. 2005. PMID: 16256419 Review.
-
From genomes to function: haloarchaea as model organisms.Microbiology (Reading). 2006 Mar;152(Pt 3):585-590. doi: 10.1099/mic.0.28504-0. Microbiology (Reading). 2006. PMID: 16514139 Review.
Cited by
-
Cellulose metabolism in halo(natrono)archaea: a comparative genomics study.Front Microbiol. 2023 Jun 1;14:1112247. doi: 10.3389/fmicb.2023.1112247. eCollection 2023. Front Microbiol. 2023. PMID: 37323904 Free PMC article.
-
Utilization of carbon catabolite repression for efficiently biotransformation of anthraquinone O-glucuronides by Streptomyces coeruleorubidus DM.Front Microbiol. 2024 Apr 16;15:1393073. doi: 10.3389/fmicb.2024.1393073. eCollection 2024. Front Microbiol. 2024. PMID: 38690368 Free PMC article.
-
Functional and Molecular Characterization of the Halomicrobium sp. IBSBa Inulosucrase.Microorganisms. 2021 Apr 2;9(4):749. doi: 10.3390/microorganisms9040749. Microorganisms. 2021. PMID: 33918392 Free PMC article.
-
Elucidating metabolic pathways through genomic analysis in highly heavy metal-resistant Halobacterium salinarum strains.Heliyon. 2024 Nov 30;10(23):e40822. doi: 10.1016/j.heliyon.2024.e40822. eCollection 2024 Dec 15. Heliyon. 2024. PMID: 39717611 Free PMC article.
-
Open Issues for Protein Function Assignment in Haloferax volcanii and Other Halophilic Archaea.Genes (Basel). 2021 Jun 24;12(7):963. doi: 10.3390/genes12070963. Genes (Basel). 2021. PMID: 34202810 Free PMC article.
References
-
- Winkenbach F, Grant BR, Bidwell RGS. 1972. The effects of nitrate, nitrite, and ammonia on photosynthetic carbon metabolism of Acetabularia chloroplast preparations compared with spinach chloroplasts and whole cells of Acetabularia and Dunaliella. Can J Bot 50:2545–2551. doi:10.1139/b72-325. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

