Small-molecule targeting of brachyury transcription factor addiction in chordoma
- PMID: 30664779
- PMCID: PMC6633917
- DOI: 10.1038/s41591-018-0312-3
Small-molecule targeting of brachyury transcription factor addiction in chordoma
Abstract
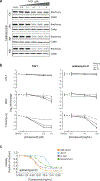
Chordoma is a primary bone cancer with no approved therapy1. The identification of therapeutic targets in this disease has been challenging due to the infrequent occurrence of clinically actionable somatic mutations in chordoma tumors2,3. Here we describe the discovery of therapeutically targetable chordoma dependencies via genome-scale CRISPR-Cas9 screening and focused small-molecule sensitivity profiling. These systematic approaches reveal that the developmental transcription factor T (brachyury; TBXT) is the top selectively essential gene in chordoma, and that transcriptional cyclin-dependent kinase (CDK) inhibitors targeting CDK7/12/13 and CDK9 potently suppress chordoma cell proliferation. In other cancer types, transcriptional CDK inhibitors have been observed to downregulate highly expressed, enhancer-associated oncogenic transcription factors4,5. In chordoma, we find that T is associated with a 1.5-Mb region containing 'super-enhancers' and is the most highly expressed super-enhancer-associated transcription factor. Notably, transcriptional CDK inhibition leads to preferential and concentration-dependent downregulation of cellular brachyury protein levels in all models tested. In vivo, CDK7/12/13-inhibitor treatment substantially reduces tumor growth. Together, these data demonstrate small-molecule targeting of brachyury transcription factor addiction in chordoma, identify a mechanism of T gene regulation that underlies this therapeutic strategy, and provide a blueprint for applying systematic genetic and chemical screening approaches to discover vulnerabilities in genomically quiet cancers.
Conflict of interest statement
COMPETING INTERESTS
T.S. is a consultant for Jnana Therapeutics. N.S.G. is equity holder and scientific advisor for Syros, Gatekeeper, Soltego, C4, Petra and Aduro companies. Syros has licensed intellectual property from Dana-Farber Cancer Institute covering THZ1. P.A. Clarke, J.B., and P.W. are current employees of The Institute of Cancer Research, which has a Rewards to Inventors scheme and has a commercial interest in the development of inhibitors of the WNT pathway, CDK8/19 and other CDKs, with intellectual property licensed to Merck and Cyclacel Pharmaceuticals. PW is a consultant for Astex Pharmaceuticals, CV6 Therapeutics, Nextechinvest, Nuevolution and Storm Therapeutics and holds equity in Chroma Therapeutics, Nextech and Storm. D.E.R. receives research funding from members of the Functional Genomics Consortium (Abbvie, Jannsen, Merck, Vir), and is a director of Addgene, Inc. W.C.H. is a consultant for Thermo Fisher, Paraxel, AjuIB, MPM Capital and KSQ Therapeutics, is a founder of KSQ Therapeutics and receives research support from Deerfield. J.E.B. is now an executive and shareholder of Novartis AG, and has been a founder and shareholder of SHAPE (acquired by Medivir), Acetylon (acquired by Celgene), Tensha (acquired by Roche), Syros, Regency and C4 Therapeutics. K.K.W. is a founder and equity holder of G1 Therapeutics and he has consulting/sponsored research agreements with AstraZeneca, Janssen, Pfizer, Array, Novartis, Merck, Takeda, Ono, Targimmune and BMS. C.Y.L. is a consultant for Jnana Therapeutics and is a shareholder of and inventor of intellectual property licensed to Syros Pharmaceuticals. J.D.K. is a founder, executive, and shareholder of Jnana Therapeutics. S.L.S. is a member of the Board of Directors of the Genomics Institute of the Novartis Research Foundation (“GNF”); a shareholder and member of the Board of Directors of Jnana Therapeutics; a shareholder of Forma Therapeutics; a shareholder of and adviser to Decibel Therapeutics; an adviser to Eisai, Inc., the Ono Pharma Foundation, and F-Prime Capital Partners; and a Novartis Faculty Scholar.
All other authors declare no competing interests.
Figures
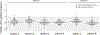












References
-
- Stacchiotti S, Sommer J & Chordoma Global Consensus Group. Building a global consensus approach to chordoma: a position paper from the medical and patient community. Lancet Oncol 16, e71–83 (2015). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

