Gut microbiota density influences host physiology and is shaped by host and microbial factors
- PMID: 30666957
- PMCID: PMC6342524
- DOI: 10.7554/eLife.40553
Gut microbiota density influences host physiology and is shaped by host and microbial factors
Abstract
To identify factors that regulate gut microbiota density and the impact of varied microbiota density on health, we assayed this fundamental ecosystem property in fecal samples across mammals, human disease, and therapeutic interventions. Physiologic features of the host (carrying capacity) and the fitness of the gut microbiota shape microbiota density. Therapeutic manipulation of microbiota density in mice altered host metabolic and immune homeostasis. In humans, gut microbiota density was reduced in Crohn's disease, ulcerative colitis, and ileal pouch-anal anastomosis. The gut microbiota in recurrent Clostridium difficile infection had lower density and reduced fitness that were restored by fecal microbiota transplantation. Understanding the interplay between microbiota and disease in terms of microbiota density, host carrying capacity, and microbiota fitness provide new insights into microbiome structure and microbiome targeted therapeutics.
Editorial note: This article has been through an editorial process in which the authors decide how to respond to the issues raised during peer review. The Reviewing Editor's assessment is that all the issues have been addressed (see decision letter).
Keywords: fecal microbiota transplantation; gut microbiome; human; human biology; infectious disease; inflammatory bowel disease; medicine; microbiology; microbiota density; mouse.
© 2019, Contijoch et al.
Conflict of interest statement
EC, GB, CY, IM, ZL, RN, SL, SH, CJ, KR, RB, ID, RH, SF, YL, NY, TL, PL, SL, IP, AG, JC, RK, SK, XQ, AC, AH, AA, JR, FF, MS, AN, PR, CR, SA, KS, LP, MS, KH, JZ, BZ, BL, HI, WS, AD, WW, BC, CD, DG, SI, AL, JM, EM, RU, SN, JN, BS, TU, PR, JG, PL, ST, JF, CB, SP, JC, JC, ES, CA, AK, BS No competing interests declared, RH, MD, JF Is a consultant for Janssen and has no other financial competing interests to declare.
Figures
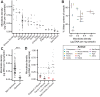




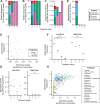
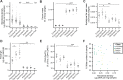




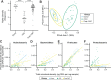
References
-
- Atarashi K, Tanoue T, Shima T, Imaoka A, Kuwahara T, Momose Y, Cheng G, Yamasaki S, Saito T, Ohba Y, Taniguchi T, Takeda K, Hori S, Ivanov II, Umesaki Y, Itoh K, Honda K. Induction of colonic regulatory T cells by indigenous clostridium species. Science. 2011;331:337–341. doi: 10.1126/science.1198469. - DOI - PMC - PubMed
-
- Battaglioli EJ, Hale VL, Chen J, Jeraldo P, Ruiz-Mojica C, Schmidt BA, Rekdal VM, Till LM, Huq L, Smits SA, Moor WJ, Jones-Hall Y, Smyrk T, Khanna S, Pardi DS, Grover M, Patel R, Chia N, Nelson H, Sonnenburg JL, Farrugia G, Kashyap PC. Clostridioides difficile uses amino acids associated with gut microbial dysbiosis in a subset of patients with diarrhea. Science Translational Medicine. 2018;10:eaam7019. doi: 10.1126/scitranslmed.aam7019. - DOI - PMC - PubMed
-
- Blandford PRS. Biology of the polecat mustela putorius: a literature review. Mammal Review. 1987;17:155–198. doi: 10.1111/j.1365-2907.1987.tb00282.x. - DOI
-
- Bongers G, Pacer ME, Geraldino TH, Chen L, He Z, Hashimoto D, Furtado GC, Ochando J, Kelley KA, Clemente JC, Merad M, van Bakel H, Lira SA. Interplay of host microbiota, genetic perturbations, and inflammation promotes local development of intestinal neoplasms in mice. The Journal of Experimental Medicine. 2014;211:457–472. doi: 10.1084/jem.20131587. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 CA161373/CA/NCI NIH HHS/United States
- DK112679/DK/NIDDK NIH HHS/United States
- S10 OD018522/OD/NIH HHS/United States
- GM007280/GM/NIGMS NIH HHS/United States
- P30 DK020541/DK/NIDDK NIH HHS/United States
- R01 GM108505/GM/NIGMS NIH HHS/United States
- T32 GM007280/GM/NIGMS NIH HHS/United States
- F31 DK112679/DK/NIDDK NIH HHS/United States
- P30 CA016087/CA/NCI NIH HHS/United States
- R01 DK110352/DK/NIDDK NIH HHS/United States
- GM108505/GM/NIGMS NIH HHS/United States
- U19 AI118610/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

