Structural Analysis of the Hanks-Type Protein Kinase YabT From Bacillus subtilis Provides New Insights in its DNA-Dependent Activation
- PMID: 30671027
- PMCID: PMC6333020
- DOI: 10.3389/fmicb.2018.03014
Structural Analysis of the Hanks-Type Protein Kinase YabT From Bacillus subtilis Provides New Insights in its DNA-Dependent Activation
Abstract
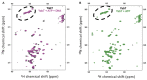
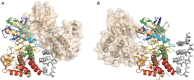
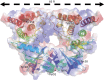
YabT is a serine/threonine kinase of the Hanks family from Bacillus subtilis, which lacks the canonical extracellular signal receptor domain but is anchored to the membrane through a C-terminal transmembrane helix. A previous study demonstrated that a basic juxtamembrane region corresponds to a DNA-binding motif essential for the activation of YabT trans-autophosphorylation. YabT is expressed during spore development and localizes to the asymmetric septum where it specifically phosphorylates essential proteins involved in genome maintenance, such as RecA, SsbA, and YabA. YabT has also been shown to phosphorylate proteins involved in protein synthesis, such as AbrB and Ef-Tu, suggesting a possible regulatory role in the progressive metabolic quiescence of the forespore. Finally, cross phosphorylations with other protein kinases implicate YabT in the regulation of numerous other cellular processes. Using an artificial protein scaffold as crystallization helper, we determined the first crystal structure of this DNA-dependent bacterial protein kinase. This allowed us to trap the active conformation of the kinase domain of YabT. Using NMR, we showed that the basic juxtamembrane region of YabT is disordered in the absence of DNA in solution, just like it is in the crystal, and that it is stabilized upon DNA binding. In comparison with its closest structural homolog, the mycobacterial kinase PknB allowed us to discuss the dimerization mode of YabT. Together with phosphorylation assays and DNA-binding experiments, this structural analysis helped us to gain new insights into the regulatory activation mechanism of YabT.
Keywords: autophosphorylation; crystallization chaperone; dimerization; regulatory mechanism; spore development.
Figures

Similar articles
-
Bacillus subtilis serine/threonine protein kinase YabT is involved in spore development via phosphorylation of a bacterial recombinase.Mol Microbiol. 2013 Jun;88(5):921-35. doi: 10.1111/mmi.12233. Epub 2013 May 2. Mol Microbiol. 2013. PMID: 23634894 Free PMC article.
-
Mass spectrometry and site-directed mutagenesis identify several autophosphorylated residues required for the activity of PrkC, a Ser/Thr kinase from Bacillus subtilis.J Mol Biol. 2003 Jul 11;330(3):459-72. doi: 10.1016/s0022-2836(03)00579-5. J Mol Biol. 2003. PMID: 12842463
-
Cross-phosphorylation of bacterial serine/threonine and tyrosine protein kinases on key regulatory residues.Front Microbiol. 2014 Sep 17;5:495. doi: 10.3389/fmicb.2014.00495. eCollection 2014. Front Microbiol. 2014. PMID: 25278935 Free PMC article.
-
Signal Sensing and Transduction by Histidine Kinases as Unveiled through Studies on a Temperature Sensor.Acc Chem Res. 2017 Jun 20;50(6):1359-1366. doi: 10.1021/acs.accounts.6b00593. Epub 2017 May 5. Acc Chem Res. 2017. PMID: 28475313 Review.
-
Impact of Serine/Threonine Protein Kinases on the Regulation of Sporulation in Bacillus subtilis.Front Microbiol. 2016 Apr 20;7:568. doi: 10.3389/fmicb.2016.00568. eCollection 2016. Front Microbiol. 2016. PMID: 27148245 Free PMC article. Review.
Cited by
-
Structural and Functional Insights into the Biofilm-Associated BceF Tyrosine Kinase Domain from Burkholderia cepacia.Biomolecules. 2021 Aug 12;11(8):1196. doi: 10.3390/biom11081196. Biomolecules. 2021. PMID: 34439861 Free PMC article.
-
Dynamics understanding of novel solvated drug molecules against emerging Burkholderia Cepacia infections in immunocompromised patients.Results Chem. 2025 May;15:None. doi: 10.1016/j.rechem.2025.102250. Results Chem. 2025. PMID: 40503072 Free PMC article.
-
Experimental measurement and computational prediction of bacterial Hanks-type Ser/Thr signaling system regulatory targets.Mol Microbiol. 2024 Aug;122(2):152-164. doi: 10.1111/mmi.15220. Epub 2024 Jan 3. Mol Microbiol. 2024. PMID: 38167835 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

