Pseudomonas bacteriocin syringacin M released upon desiccation suppresses the growth of sensitive bacteria in plant necrotic lesions
- PMID: 30672132
- PMCID: PMC6922522
- DOI: 10.1111/1751-7915.13367
Pseudomonas bacteriocin syringacin M released upon desiccation suppresses the growth of sensitive bacteria in plant necrotic lesions
Abstract
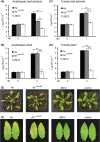
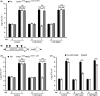
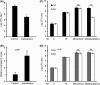
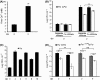
Bacteriocins are regarded as important factors mediating microbial interactions, but their exact role in community ecology largely remains to be elucidated. Here, we report the characterization of a mutant strain, derived from Pseudomonas syringae pv. tomato DC3000 (Pst), that was incapable of growing in plant extracts and causing disease. Results showed that deficiency in a previously unannotated gene saxE led to the sensitivity of the mutant to Ca2+ in leaf extracts. Transposon insertions in the bacteriocin gene syrM, adjacent to saxE, fully rescued the bacterial virulence and growth of the ΔsaxE mutant in plant extracts, indicating that syrM-saxE encode a pair of bacteriocin immunity proteins in Pst. To investigate whether the syrM-saxE system conferred any advantage to Pst in competition with other SyrM-sensitive pathovars, we compared the growth of a SyrM-sensitive strain co-inoculated with Pst strains with or without the syrM gene and observed a significant syrM-dependent growth reduction of the sensitive bacteria on plate and in lesion tissues upon desiccation-rehydration treatment. These findings reveal an important biological role of SyrM-like bacteriocins and help to understand the complex strategies used by P. syringae in adaptation to the phyllosphere niche in the context of plant disease.
© 2019 The Authors. Microbial Biotechnology published by John Wiley & Sons Ltd and Society for Applied Microbiology.
Conflict of interest statement
None declared.
Figures


Similar articles
-
The plant host environment influences competitive interactions between bacterial pathogens.Environ Microbiol Rep. 2022 Oct;14(5):785-794. doi: 10.1111/1758-2229.13103. Epub 2022 Jun 14. Environ Microbiol Rep. 2022. PMID: 35700743 Free PMC article.
-
Independent Co-Option of a Tailed Bacteriophage into a Killing Complex in Pseudomonas.mBio. 2015 Aug 11;6(4):e00452. doi: 10.1128/mBio.00452-15. mBio. 2015. PMID: 26265717 Free PMC article.
-
IDL6-HAE/HSL2 impacts pectin degradation and resistance to Pseudomonas syringae pv tomato DC3000 in Arabidopsis leaves.Plant J. 2017 Jan;89(2):250-263. doi: 10.1111/tpj.13380. Epub 2017 Jan 16. Plant J. 2017. PMID: 27618493
-
Pseudomonas syringae pv. tomato DC3000: a model pathogen for probing disease susceptibility and hormone signaling in plants.Annu Rev Phytopathol. 2013;51:473-98. doi: 10.1146/annurev-phyto-082712-102321. Epub 2013 May 31. Annu Rev Phytopathol. 2013. PMID: 23725467 Review.
-
Pseudomonas savastanoi pv. savastanoi: some like it knot.Mol Plant Pathol. 2012 Dec;13(9):998-1009. doi: 10.1111/j.1364-3703.2012.00816.x. Epub 2012 Jul 17. Mol Plant Pathol. 2012. PMID: 22805238 Free PMC article. Review.
Cited by
-
Bacteriocins Targeting Gram-Negative Phytopathogenic Bacteria: Plantibiotics of the Future.Front Microbiol. 2020 Sep 18;11:575981. doi: 10.3389/fmicb.2020.575981. eCollection 2020. Front Microbiol. 2020. PMID: 33042091 Free PMC article. Review.
-
The plant host environment influences competitive interactions between bacterial pathogens.Environ Microbiol Rep. 2022 Oct;14(5):785-794. doi: 10.1111/1758-2229.13103. Epub 2022 Jun 14. Environ Microbiol Rep. 2022. PMID: 35700743 Free PMC article.
-
Pseudomonas syringae Type III Secretion Protein HrpP Manipulates Plant Immunity To Promote Infection.Microbiol Spectr. 2023 Jun 15;11(3):e0514822. doi: 10.1128/spectrum.05148-22. Epub 2023 Apr 17. Microbiol Spectr. 2023. PMID: 37067445 Free PMC article.
-
Bacteriocins: potentials and prospects in health and agrifood systems.Arch Microbiol. 2024 Apr 25;206(5):233. doi: 10.1007/s00203-024-03948-y. Arch Microbiol. 2024. PMID: 38662051 Free PMC article. Review.
-
A putative nuclear copper chaperone promotes plant immunity in Arabidopsis.J Exp Bot. 2020 Oct 22;71(20):6684-6696. doi: 10.1093/jxb/eraa401. J Exp Bot. 2020. PMID: 32865553 Free PMC article.
References
-
- And, S.S.H. , and Upper, C.D. (1983) Ecology and epidemiology of foliar bacterial plant pathogens. Annu Rev Phytopathol 21: 243–270.
-
- Barreteau, H. , El Ghachi, M. , Barnéoud‐Arnoulet, A. , Sacco, E. , Touzé, T. , Duché, D. , et al (2012) Characterization of colicin M and its orthologs targeting bacterial cell wall peptidoglycan biosynthesis. Microb Drug Resist 18: 222–229. - PubMed
-
- Beattie, G.A. (2011) Water relations in the interaction of foliar bacterial pathogens with plants. Annu Rev Phytopathol 49: 533–555. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

