Structural insights into the activation of metabotropic glutamate receptors
- PMID: 30675062
- PMCID: PMC6709600
- DOI: 10.1038/s41586-019-0881-4
Structural insights into the activation of metabotropic glutamate receptors
Erratum in
-
Author Correction: Structural insights into the activation of metabotropic glutamate receptors.Nature. 2019 Mar;567(7747):E10. doi: 10.1038/s41586-019-0983-z. Nature. 2019. PMID: 30814739
Abstract
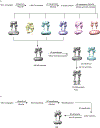
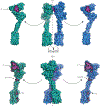
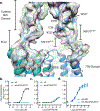
Metabotropic glutamate receptors are family C G-protein-coupled receptors. They form obligate dimers and possess extracellular ligand-binding Venus flytrap domains, which are linked by cysteine-rich domains to their 7-transmembrane domains. Spectroscopic studies show that signalling is a dynamic process, in which large-scale conformational changes underlie the transmission of signals from the extracellular Venus flytraps to the G protein-coupling domains-the 7-transmembrane domains-in the membrane. Here, using a combination of X-ray crystallography, cryo-electron microscopy and signalling studies, we present a structural framework for the activation mechanism of metabotropic glutamate receptor subtype 5. Our results show that agonist binding at the Venus flytraps leads to a compaction of the intersubunit dimer interface, thereby bringing the cysteine-rich domains into close proximity. Interactions between the cysteine-rich domains and the second extracellular loops of the receptor enable the rigid-body repositioning of the 7-transmembrane domains, which come into contact with each other to initiate signalling.
Conflict of interest statement
The authors declare one competing interest: Brian Kobilka and Tong Sun Kobilka are founders of and consultants for ConfometRx, Inc. Dan Feng and Bingfa Sun are employees of ConfometRx, Inc., and Matthew Chu was an employee of ConfometRx, Inc. Readers are welcome to comment on the online version of the paper.
Figures



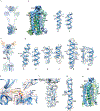
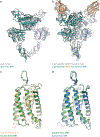
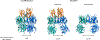
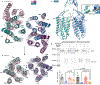

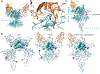
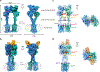
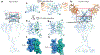

Comment in
-
How an activation signal is transmitted through an excitatory receptor.Nature. 2019 Feb;566(7742):42-43. doi: 10.1038/d41586-018-07885-x. Nature. 2019. PMID: 30710123 No abstract available.
Similar articles
-
[Recent advances in the structural biology of the class C G protein-coupled receptors: The metabotropic Glutamate receptor 5].Biol Aujourdhui. 2021;215(3-4):85-94. doi: 10.1051/jbio/2021013. Epub 2022 Mar 11. Biol Aujourdhui. 2021. PMID: 35275053 Review. French.
-
Agonists and allosteric modulators promote signaling from different metabotropic glutamate receptor 5 conformations.Cell Rep. 2021 Aug 31;36(9):109648. doi: 10.1016/j.celrep.2021.109648. Cell Rep. 2021. PMID: 34469715 Free PMC article.
-
Conformational diversity in class C GPCR positive allosteric modulation.Nat Commun. 2025 Jan 13;16(1):619. doi: 10.1038/s41467-024-55439-9. Nat Commun. 2025. PMID: 39805839 Free PMC article.
-
Stepwise activation of a metabotropic glutamate receptor.Nature. 2024 May;629(8013):951-956. doi: 10.1038/s41586-024-07327-x. Epub 2024 Apr 17. Nature. 2024. PMID: 38632403 Free PMC article.
-
Structure-based discovery and development of metabotropic glutamate receptor 5 negative allosteric modulators.Adv Pharmacol. 2020;88:35-58. doi: 10.1016/bs.apha.2020.03.001. Epub 2020 Mar 25. Adv Pharmacol. 2020. PMID: 32416871 Review.
Cited by
-
Current Progress in Understanding the Structure and Function of Sweet Taste Receptor.J Mol Neurosci. 2021 Feb;71(2):234-244. doi: 10.1007/s12031-020-01642-4. Epub 2020 Jun 30. J Mol Neurosci. 2021. PMID: 32607758 Review.
-
Electrostatic Surface Potential as a Key Parameter in Virus Transmission and Evolution: How to Manage Future Virus Pandemics in the Post-COVID-19 Era.Viruses. 2023 Jan 19;15(2):284. doi: 10.3390/v15020284. Viruses. 2023. PMID: 36851498 Free PMC article. Review.
-
Oligomerization of yeast α-factor receptor detected by fluorescent energy transfer between ligands.Biophys J. 2021 Nov 16;120(22):5090-5106. doi: 10.1016/j.bpj.2021.10.005. Epub 2021 Oct 8. Biophys J. 2021. PMID: 34627767 Free PMC article.
-
Structural insights into dimerization and activation of the mGlu2-mGlu3 and mGlu2-mGlu4 heterodimers.Cell Res. 2023 Oct;33(10):762-774. doi: 10.1038/s41422-023-00830-2. Epub 2023 Jun 8. Cell Res. 2023. PMID: 37286794 Free PMC article.
-
Pharmacology of the Umami Taste Receptor.Handb Exp Pharmacol. 2022;275:109-136. doi: 10.1007/164_2021_439. Handb Exp Pharmacol. 2022. PMID: 33580387
References
-
- Pin J-P & Bettler B Organization and functions of mGlu and GABA. Nature 540, 60–68 (2016). - PubMed
-
- Kunishima N et al. Structural basis of glutamate recognition by a dimeric metabotropic glutamate receptor. Nature 407, 971–977 (2000). - PubMed
-
- Kniazeff J et al. Closed state of both binding domains of homodimeric mGlu receptors is required for full activity. Nat. Struct. Mol. Biol 11, 706–713 (2004). - PubMed
-
- Hlavackova V et al. Sequential inter- and intrasubunit rearrangements during activation of dimeric metabotropic glutamate receptor 1. Sci Signal 5, ra59–ra59 (2012). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

