Increased predominance of HIV-1 CRF01_AE and its recombinants in the Philippines
- PMID: 30676308
- PMCID: PMC7011713
- DOI: 10.1099/jgv.0.001198
Increased predominance of HIV-1 CRF01_AE and its recombinants in the Philippines
Abstract
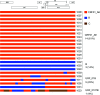
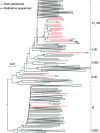
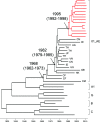
The growth rate of new HIV infections in the Philippines was the fastest of any countries in the Asia-Pacific region between 2010 and 2016. To date, HIV-1 subtyping results in the Philippines have been determined by characterizing only partial viral genome sequences. It is not known whether recombination occurs in the majority of unsequenced genome regions. Near-full-length genome (NFLG) sequences were obtained by amplifying two overlapping half genomes from plasma samples collected between 2015 and 2017 from 23 newly diagnosed infected individuals in the Philippines. Phylogenetic analysis showed that the newly characterized sequences were CRF01_AE (14), subtype B (3), CRF01/B recombinants (5) and a CRF01/CRF07/B recombinant (1). All 14 CRF01_AE formed a tight cluster, suggesting that they were derived from a single introduction. The time to the most recent common ancestor (tMRCA) for CRF01_AE in the Philippines was 1995 (1992-1998), about 10-15 years later than that of CRF01_AE in China and Thailand. All five CRF01/B recombinants showed distinct recombination patterns, suggesting ongoing recombination between the two predominant circulating viruses. The identification of partial CRF07_BC sequences in one CRF01/CRF07/B recombinant, not reported previously in the Philippines, indicated that CRF07_BC may have been recently introduced into that country from China, where CRF07_BC is prevalent. Our results show that the major epidemic strains may have shifted to an increased predominance of CRF01_AE and its recombinants, and that other genotypes such as CRF07_BC may have been introduced into the Philippines.
Keywords: DRMs; HIV-1; NFLG; Philippines; subtype; tMRCA.
Conflict of interest statement
The authors declare no completing interests.
Figures



References
-
- Joint United Nations Programme on HIV/AIDS (UNAIDS) UNAIDS DATA 2017. Geneva, Switzerland: 2017. www.unaids.org/sites/default/files/media_asset/20170720_Data_book_2017_e...
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

