Unraveling the cellular origin and clinical prognostic markers of infant B-cell acute lymphoblastic leukemia using genome-wide analysis
- PMID: 30679323
- PMCID: PMC6545849
- DOI: 10.3324/haematol.2018.206375
Unraveling the cellular origin and clinical prognostic markers of infant B-cell acute lymphoblastic leukemia using genome-wide analysis
Abstract
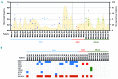
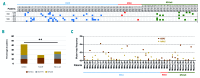
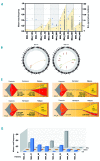
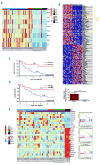
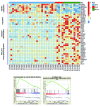
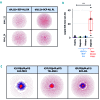
B-cell acute lymphoblastic leukemia is the commonest childhood cancer. In infants, B-cell acute lymphoblastic leukemia remains fatal, especially in patients with t(4;11), present in ~80% of cases. The pathogenesis of t(4;11)/KMT2A-AFF1+ (MLL-AF4+) infant B-cell acute lymphoblastic leukemia remains difficult to model, and the pathogenic contribution in cancer of the reciprocal fusions resulting from derivative translocated-chromosomes remains obscure. Here, "multi-layered" genome-wide analyses and validation were performed on a total of 124 de novo cases of infant B-cell acute lymphoblastic leukemia uniformly diagnosed and treated according to the Interfant 99/06 protocol. These patients showed the most silent mutational landscape reported so far for any sequenced pediatric cancer. Recurrent mutations were exclusively found in K-RAS and N-RAS, were subclonal and were frequently lost at relapse, despite a larger number of non-recurrent/non-silent mutations. Unlike non-MLL-rearranged B-cell acute lymphoblastic leukemias, B-cell receptor repertoire analysis revealed minor, non-expanded B-cell clones in t(4;11)+ infant B-cell acute lymphoblastic leukemia, and RNA-sequencing showed transcriptomic similarities between t(4;11)+ infant B-cell acute lymphoblastic leukemias and the most immature human fetal liver hematopoietic stem and progenitor cells, confirming a "pre-VDJ" fetal cellular origin for both t(4;11) and RAS mut The reciprocal fusion AF4-MLL was expressed in only 45% (19/43) of the t(4;11)+ patients, and HOXA cluster genes are exclusively expressed in AF4-MLL-expressing patients. Importantly, AF4-MLL/HOXA-expressing patients had a significantly better 4-year event-free survival (62.4% vs 11.7%, P=0.001), and overall survival (73.7 vs 25.2%, P=0.016). AF4-MLL expression retained its prognostic significance when analyzed in a Cox model adjusting for risk stratification according to the Interfant-06 protocol based on age at diagnosis, white blood cell count and response to prednisone. This study has clinical implications for disease outcome and diagnostic risk-stratification of t(4;11)+ infant B-cell acute lymphoblastic leukemia.
Copyright© 2019 Ferrata Storti Foundation.
Figures

Comment in
-
Another piece of the puzzle added to understand t(4;11) leukemia better.Haematologica. 2019 Jun;104(6):1098-1100. doi: 10.3324/haematol.2018.213397. Haematologica. 2019. PMID: 31152087 Free PMC article. No abstract available.
References
-
- Marschalek R. Mechanisms of leukemogenesis by MLL fusion proteins. Br J Haematol. 2011;152(2):141–154. - PubMed

