The Short Chain Fatty Acid Butyrate Imprints an Antimicrobial Program in Macrophages
- PMID: 30683619
- PMCID: PMC6382411
- DOI: 10.1016/j.immuni.2018.12.018
The Short Chain Fatty Acid Butyrate Imprints an Antimicrobial Program in Macrophages
Abstract
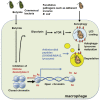
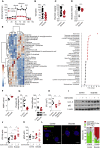
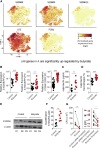
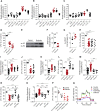
Host microbial cross-talk is essential to maintain intestinal homeostasis. However, maladaptation of this response through microbial dysbiosis or defective host defense toward invasive intestinal bacteria can result in chronic inflammation. We have shown that macrophages differentiated in the presence of the bacterial metabolite butyrate display enhanced antimicrobial activity. Butyrate-induced antimicrobial activity was associated with a shift in macrophage metabolism, a reduction in mTOR kinase activity, increased LC3-associated host defense and anti-microbial peptide production in the absence of an increased inflammatory cytokine response. Butyrate drove this monocyte to macrophage differentiation program through histone deacetylase 3 (HDAC3) inhibition. Administration of butyrate induced antimicrobial activity in intestinal macrophages in vivo and increased resistance to enteropathogens. Our data suggest that (1) increased intestinal butyrate might represent a strategy to bolster host defense without tissue damaging inflammation and (2) that pharmacological HDAC3 inhibition might drive selective macrophage functions toward antimicrobial host defense.
Copyright © 2018 The Authors. Published by Elsevier Inc. All rights reserved.
Figures


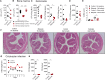
Comment in
-
Butyrate boosts microbicidal macrophages.Nat Rev Immunol. 2019 Mar;19(3):135. doi: 10.1038/s41577-019-0132-9. Nat Rev Immunol. 2019. PMID: 30723307 No abstract available.
-
Butyrate Makes Macrophages "Go Nuclear" against Bacterial Pathogens.Immunity. 2019 Feb 19;50(2):275-278. doi: 10.1016/j.immuni.2019.01.015. Immunity. 2019. PMID: 30784572
References
-
- Ananthakrishnan A.N., Bernstein C.N., Iliopoulos D., Macpherson A., Neurath M.F., Ali R.A.R., Vavricka S.R., Fiocchi C. Environmental triggers in IBD: a review of progress and evidence. Nat. Rev. Gastroenterol. Hepatol. 2018;15:39–49. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

