An integrated transcriptional analysis of the developing human retina
- PMID: 30696714
- PMCID: PMC6361134
- DOI: 10.1242/dev.169474
An integrated transcriptional analysis of the developing human retina
Abstract
The scarcity of embryonic/foetal material as a resource for direct study means that there is still limited understanding of human retina development. Here, we present an integrated transcriptome analysis combined with immunohistochemistry in human eye and retinal samples from 4 to 19 post-conception weeks. This analysis reveals three developmental windows with specific gene expression patterns that informed the sequential emergence of retinal cell types and enabled identification of stage-specific cellular and biological processes, and transcriptional regulators. Each stage is characterised by a specific set of alternatively spliced transcripts that code for proteins involved in the formation of the photoreceptor connecting cilium, pre-mRNA splicing and epigenetic modifiers. Importantly, our data show that the transition from foetal to adult retina is characterised by a large increase in the percentage of mutually exclusive exons that code for proteins involved in photoreceptor maintenance. The circular RNA population is also defined and shown to increase during retinal development. Collectively, these data increase our understanding of human retinal development and the pre-mRNA splicing process, and help to identify new candidate disease genes.
Keywords: Alternative splicing; Circular RNAs; Developing human retina; Immunohistochemistry; Retinal cells; Transcriptome.
© 2019. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interestsThe authors declare no competing or financial interests.
Figures



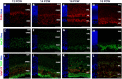
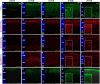
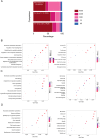


References
-
- Aldiri I., Xu B., Wang L., Chen X., Hiler D., Griffiths L., Valentine M., Shirinifard A., Thiagarajan S., Sablauer A. et al. (2017). The dynamic epigenetic landscape of the retina during development, reprogramming, and tumorigenesis. Neuron 94, 550-568. 10.1016/j.neuron.2017.04.022 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- MC_PC_15030/MRC_/Medical Research Council/United Kingdom
- MC_UP_1501/2/MRC_/Medical Research Council/United Kingdom
- MR/M008886/1/MRC_/Medical Research Council/United Kingdom
- MR/N015037/1/MRC_/Medical Research Council/United Kingdom
- 614620/ERC_/European Research Council/International
- MR/R006237/1/MRC_/Medical Research Council/United Kingdom
- NC/C016106/1/NC3RS_/National Centre for the Replacement, Refinement and Reduction of Animals in Research/United Kingdom
- 099175/Z/12/Z/WT_/Wellcome Trust/United Kingdom
- BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- MC_UU_00015/9/MRC_/Medical Research Council/United Kingdom
- WT_/Wellcome Trust/United Kingdom
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

