Detection of RNA-DNA binding sites in long noncoding RNAs
- PMID: 30698727
- PMCID: PMC6451187
- DOI: 10.1093/nar/gkz037
Detection of RNA-DNA binding sites in long noncoding RNAs
Abstract
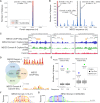
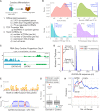
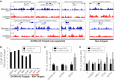
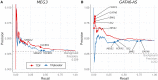
Long non-coding RNAs (lncRNAs) can act as scaffolds that promote the interaction of proteins, RNA, and DNA. There is increasing evidence of sequence-specific interactions of lncRNAs with DNA via triple-helix (triplex) formation. This process allows lncRNAs to recruit protein complexes to specific genomic regions and regulate gene expression. Here we propose a computational method called Triplex Domain Finder (TDF) to detect triplexes and characterize DNA-binding domains and DNA targets statistically. Case studies showed that this approach can detect the known domains of lncRNAs Fendrr, HOTAIR and MEG3. Moreover, we validated a novel DNA-binding domain in MEG3 by a genome-wide sequencing method. We used TDF to perform a systematic analysis of the triplex-forming potential of lncRNAs relevant to human cardiac differentiation. We demonstrated that the lncRNA with the highest triplex-forming potential, GATA6-AS, forms triple helices in the promoter of genes relevant to cardiac development. Moreover, down-regulation of GATA6-AS impairs GATA6 expression and cardiac development. These data indicate the unique ability of our computational tool to identify novel triplex-forming lncRNAs and their target genes.
© The Author(s) 2019. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

Similar articles
-
Practical Guidance in Genome-Wide RNA:DNA Triple Helix Prediction.Int J Mol Sci. 2020 Jan 28;21(3):830. doi: 10.3390/ijms21030830. Int J Mol Sci. 2020. PMID: 32012884 Free PMC article.
-
Genome-wide computational analysis of potential long noncoding RNA mediated DNA:DNA:RNA triplexes in the human genome.J Transl Med. 2017 Sep 2;15(1):186. doi: 10.1186/s12967-017-1282-9. J Transl Med. 2017. PMID: 28865451 Free PMC article.
-
Super-lncRNAs: identification of lncRNAs that target super-enhancers via RNA:DNA:DNA triplex formation.RNA. 2017 Nov;23(11):1729-1742. doi: 10.1261/rna.061317.117. Epub 2017 Aug 24. RNA. 2017. PMID: 28839111 Free PMC article.
-
RNA-DNA Triplex Formation by Long Noncoding RNAs.Cell Chem Biol. 2016 Nov 17;23(11):1325-1333. doi: 10.1016/j.chembiol.2016.09.011. Epub 2016 Oct 20. Cell Chem Biol. 2016. PMID: 27773629 Review.
-
Biophysical Investigation of RNA ⋅ DNA : DNA Triple Helix and RNA : DNA Heteroduplex Formation by the lncRNAs MEG3 and Fendrr.Chembiochem. 2024 May 17;25(10):e202400049. doi: 10.1002/cbic.202400049. Epub 2024 Apr 29. Chembiochem. 2024. PMID: 38456652 Review.
Cited by
-
HIF1α-AS1 is a DNA:DNA:RNA triplex-forming lncRNA interacting with the HUSH complex.Nat Commun. 2022 Nov 2;13(1):6563. doi: 10.1038/s41467-022-34252-2. Nat Commun. 2022. PMID: 36323673 Free PMC article.
-
Long Non-coding RNA PVT1 as a Prognostic and Therapeutic Target in Pediatric Cancer.Front Oncol. 2019 Nov 6;9:1173. doi: 10.3389/fonc.2019.01173. eCollection 2019. Front Oncol. 2019. PMID: 31781490 Free PMC article. Review.
-
Detecting TAD-like domains from RNA-associated interactions.Nucleic Acids Res. 2022 Aug 26;50(15):e88. doi: 10.1093/nar/gkac422. Nucleic Acids Res. 2022. PMID: 35639502 Free PMC article.
-
Fasim-LongTarget enables fast and accurate genome-wide lncRNA/DNA binding prediction.Comput Struct Biotechnol J. 2022 Jun 18;20:3347-3350. doi: 10.1016/j.csbj.2022.06.017. eCollection 2022. Comput Struct Biotechnol J. 2022. PMID: 35832611 Free PMC article.
-
Nucleus size and DNA accessibility are linked to the regulation of paraspeckle formation in cellular differentiation.BMC Biol. 2020 Apr 22;18(1):42. doi: 10.1186/s12915-020-00770-y. BMC Biol. 2020. PMID: 32321486 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

