Metabolomic and Transcriptomic Analyses of Escherichia coli for Efficient Fermentation of L-Fucose
- PMID: 30699916
- PMCID: PMC6410053
- DOI: 10.3390/md17020082
Metabolomic and Transcriptomic Analyses of Escherichia coli for Efficient Fermentation of L-Fucose
Abstract
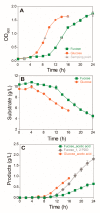
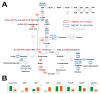
L-Fucose, one of the major monomeric sugars in brown algae, possesses high potential for use in the large-scale production of bio-based products. Although fucose catabolic pathways have been enzymatically evaluated, the effects of fucose as a carbon source on intracellular metabolism in industrial microorganisms such as Escherichia coli are still not identified. To elucidate the effects of fucose on cellular metabolism and to find clues for efficient conversion of fucose into bio-based products, comparative metabolomic and transcriptomic analyses were performed on E. coli on L-fucose and on D-glucose as a control. When fucose was the carbon source for E. coli, integration of the two omics analyses revealed that excess gluconeogenesis and quorum sensing led to severe depletion of ATP, resulting in accumulation and export of fucose extracellularly. Therefore, metabolic engineering and optimization are needed for E. coil to more efficiently ferment fucose. This is the first multi-omics study investigating the effects of fucose on cellular metabolism in E. coli. These omics data and their biological interpretation could be used to assist metabolic engineering of E. coli producing bio-based products using fucose-containing brown macroalgae.
Keywords: Escherichia coli; brown macroalgae; fermentation; fucose; metabolomics; transcriptomics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
Functional Analysis of Deoxyhexose Sugar Utilization in Escherichia coli Reveals Fermentative Metabolism under Aerobic Conditions.Appl Environ Microbiol. 2021 Jul 27;87(16):e0071921. doi: 10.1128/AEM.00719-21. Epub 2021 Jul 27. Appl Environ Microbiol. 2021. PMID: 34047632 Free PMC article.
-
Optimization of l-Fucose Biosynthesis in Escherichia coli through Pathway Engineering and Mixed Carbon Source Strategy.J Agric Food Chem. 2025 Mar 12;73(10):6102-6112. doi: 10.1021/acs.jafc.4c12544. Epub 2025 Mar 3. J Agric Food Chem. 2025. PMID: 40029204
-
L-Fucose production by engineered Escherichia coli.Biotechnol Bioeng. 2019 Apr;116(4):904-911. doi: 10.1002/bit.26907. Epub 2019 Feb 4. Biotechnol Bioeng. 2019. PMID: 30597526
-
[Advances in the biosynthesis of L-homoserine and its derivatives by metabolic engineering of Escherichia coli].Sheng Wu Gong Cheng Xue Bao. 2022 Dec 25;38(12):4385-4402. doi: 10.13345/j.cjb.220377. Sheng Wu Gong Cheng Xue Bao. 2022. PMID: 36593184 Review. Chinese.
-
Metabolic engineering of Escherichia coli for efficient production of l-arginine.Adv Appl Microbiol. 2023;122:127-150. doi: 10.1016/bs.aambs.2022.11.002. Epub 2022 Dec 20. Adv Appl Microbiol. 2023. PMID: 37085192 Review.
Cited by
-
Strain engineering and metabolic flux analysis of a probiotic yeast Saccharomyces boulardii for metabolizing L-fucose, a mammalian mucin component.Microb Cell Fact. 2022 Oct 7;21(1):204. doi: 10.1186/s12934-022-01926-x. Microb Cell Fact. 2022. PMID: 36207743 Free PMC article.
-
Environmental modulators of algae-bacteria interactions at scale.Cell Syst. 2024 Sep 18;15(9):838-853.e13. doi: 10.1016/j.cels.2024.08.002. Epub 2024 Sep 4. Cell Syst. 2024. PMID: 39236710
-
L-Fucose is involved in human-gut microbiome interactions.Appl Microbiol Biotechnol. 2023 Jun;107(12):3869-3875. doi: 10.1007/s00253-023-12527-y. Epub 2023 May 6. Appl Microbiol Biotechnol. 2023. PMID: 37148338 Review.
-
Exercise postconditioning reduces ischemic injury via suppression of cerebral gluconeogenesis in rats.Brain Behav. 2023 Jan;13(1):e2805. doi: 10.1002/brb3.2805. Epub 2022 Nov 30. Brain Behav. 2023. PMID: 36448290 Free PMC article.
-
Fucose ameliorates the proinflammatory property of Fusobacterium nucleatum in colitis via altering its metabolism.Front Cell Infect Microbiol. 2023 May 1;13:1190602. doi: 10.3389/fcimb.2023.1190602. eCollection 2023. Front Cell Infect Microbiol. 2023. PMID: 37197204 Free PMC article.
References
-
- Alonso D.M., Bond J.Q., Dumesic J.A. Catalytic conversion of biomass to biofuels. Green Chem. 2010;12:1493–1513. doi: 10.1039/c004654j. - DOI
-
- Andriamanantoanina H., Rinaudo M. Characterization of the alginates from five madagascan brown algae. Carbohyd. Polym. 2010;82:555–560. doi: 10.1016/j.carbpol.2010.05.002. - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

