Structural Asymmetry and Kinetic Limping of Single Rotary F-ATP Synthases
- PMID: 30704145
- PMCID: PMC6384691
- DOI: 10.3390/molecules24030504
Structural Asymmetry and Kinetic Limping of Single Rotary F-ATP Synthases
Abstract
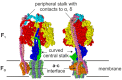
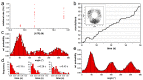
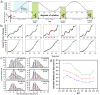
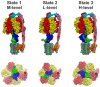
F-ATP synthases use proton flow through the FO domain to synthesize ATP in the F₁ domain. In Escherichia coli, the enzyme consists of rotor subunits γεc10 and stator subunits (αβ)₃δab₂. Subunits c10 or (αβ)₃ alone are rotationally symmetric. However, symmetry is broken by the b₂ homodimer, which together with subunit δa, forms a single eccentric stalk connecting the membrane embedded FO domain with the soluble F₁ domain, and the central rotating and curved stalk composed of subunit γε. Although each of the three catalytic binding sites in (αβ)₃ catalyzes the same set of partial reactions in the time average, they might not be fully equivalent at any moment, because the structural symmetry is broken by contact with b₂δ in F₁ and with b₂a in FO. We monitored the enzyme's rotary progression during ATP hydrolysis by three single-molecule techniques: fluorescence video-microscopy with attached actin filaments, Förster resonance energy transfer between pairs of fluorescence probes, and a polarization assay using gold nanorods. We found that one dwell in the three-stepped rotary progression lasting longer than the other two by a factor of up to 1.6. This effect of the structural asymmetry is small due to the internal elastic coupling.
Keywords: Escherichia coli; FOF1 ATP synthase; cryo-EM structure; elasticity; single-molecule fluorescence; subunit rotation; symmetry.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Rotor/Stator interactions of the epsilon subunit in Escherichia coli ATP synthase and implications for enzyme regulation.J Biol Chem. 2004 Aug 20;279(34):35616-21. doi: 10.1074/jbc.M405012200. Epub 2004 Jun 15. J Biol Chem. 2004. PMID: 15199054
-
The regulatory subunit ε in Escherichia coli FOF1-ATP synthase.Biochim Biophys Acta Bioenerg. 2018 Sep;1859(9):775-788. doi: 10.1016/j.bbabio.2018.06.013. Epub 2018 Jun 20. Biochim Biophys Acta Bioenerg. 2018. PMID: 29932911 Free PMC article. Review.
-
Three-color Förster resonance energy transfer within single F₀F₁-ATP synthases: monitoring elastic deformations of the rotary double motor in real time.J Biomed Opt. 2012 Jan;17(1):011004. doi: 10.1117/1.JBO.17.1.011004. J Biomed Opt. 2012. PMID: 22352638
-
Cryo-EM structures provide insight into how E. coli F1Fo ATP synthase accommodates symmetry mismatch.Nat Commun. 2020 May 26;11(1):2615. doi: 10.1038/s41467-020-16387-2. Nat Commun. 2020. PMID: 32457314 Free PMC article.
-
Rotating proton pumping ATPases: subunit/subunit interactions and thermodynamics.IUBMB Life. 2013 Mar;65(3):247-54. doi: 10.1002/iub.1134. IUBMB Life. 2013. PMID: 23441040 Review.
Cited by
-
F1-ATPase Rotary Mechanism: Interpreting Results of Diverse Experimental Modes With an Elastic Coupling Theory.Front Microbiol. 2022 Apr 22;13:861855. doi: 10.3389/fmicb.2022.861855. eCollection 2022. Front Microbiol. 2022. PMID: 35531282 Free PMC article. Review.
-
The nucleotide binding affinities of two critical conformations of Escherichia coli ATP synthase.Arch Biochem Biophys. 2021 Aug 15;707:108899. doi: 10.1016/j.abb.2021.108899. Epub 2021 May 12. Arch Biochem Biophys. 2021. PMID: 33991499 Free PMC article.
-
Direct observation of stepping rotation of V-ATPase reveals rigid component in coupling between Vo and V1 motors.Proc Natl Acad Sci U S A. 2022 Oct 18;119(42):e2210204119. doi: 10.1073/pnas.2210204119. Epub 2022 Oct 10. Proc Natl Acad Sci U S A. 2022. PMID: 36215468 Free PMC article.
-
Cryo-EM reveals distinct conformations of E. coli ATP synthase on exposure to ATP.Elife. 2019 Mar 26;8:e43864. doi: 10.7554/eLife.43864. Elife. 2019. PMID: 30912741 Free PMC article.
-
pH-dependent 11° F1FO ATP synthase sub-steps reveal insight into the FO torque generating mechanism.Elife. 2021 Dec 31;10:e70016. doi: 10.7554/eLife.70016. Elife. 2021. PMID: 34970963 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

