Genome analysis of Salmonella enterica subsp. diarizonae isolates from invasive human infections reveals enrichment of virulence-related functions in lineage ST1256
- PMID: 30704413
- PMCID: PMC6357384
- DOI: 10.1186/s12864-018-5352-z
Genome analysis of Salmonella enterica subsp. diarizonae isolates from invasive human infections reveals enrichment of virulence-related functions in lineage ST1256
Erratum in
-
Correction to: Genome analysis of Salmonella enterica subsp. diarizonae isolates from invasive human infections reveals enrichment of virulence-related functions in lineage ST1256.BMC Genomics. 2020 May 26;21(1):373. doi: 10.1186/s12864-020-06781-x. BMC Genomics. 2020. PMID: 32456693 Free PMC article.
Abstract
Background: Salmonella enterica subsp. diarizonae (IIIb) is frequently isolated from the environment, cold-blooded reptiles, sheep and humans; however only a few studies describe the isolation of this subspecies from invasive human infections. The factors contributing to this unusual behavior are currently unknown.
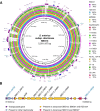
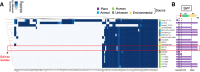
Results: We report here the genome features of two diarizonae strains, SBO13 and SBO27, isolated from endocervical tissue collected post-abortion and from cerebrospinal fluid of a newborn child, respectively, in the city of Santa Cruz, Bolivia. Although isolated six years apart, SBO27 in 2008 and SBO13 in 2014, both strains belong to the same sequence type 1256 (ST1256) and show a high degree of genome conservation sharing more than 99% of their genes, including the conservation of a ~ 10 kb plasmid. A prominent feature of the two genomes is the presence of 24 genomic islands (GIs), in addition to 10 complete Salmonella pathogenicity islands (SPI) and fragments of SPI-7, a pathogenicity island first reported in the human-adapted serovar Typhi. Some of the GIs identified in SBO13 and SBO27 harbor genes putatively encoding auto-transporters involved in adhesion, lipopolysaccharide modifying enzymes, putative toxins, pili-related proteins, efflux pumps, and several putative membrane cation transport related-genes, among others. These two Bolivian isolates also share genes encoding the type-III secretion system effector proteins SseK2, SseK3 and SlrP with other diarizonae sequence types (ST) mainly-associated with infections in humans. The sseK2, sseK3 and slrP genes were either absent or showing frameshift mutations in a significant proportion of genomes from environmental diarizonae isolates.
Conclusions: The comparative genomic study of two diarizonae strains isolated in Bolivia from human patients uncovered the presence of many genes putatively related to virulence. The statistically-significant acquisition of a unique combination of these functions by diarizonae strains isolated from humans may have impacted the ability of these isolates to successfully infect the human host.
Keywords: Comparative genomics; Invasive human infections; Salmonella enterica; Subspecies diarizonae; Type-III effectors; Virulence genes.
Conflict of interest statement
Ethics approval and consent to participate
Human endocervical tissue and cerebrospinal fluid in which
Consent for publication
Not applicable.
Competing interests
The authors declare they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures



References
-
- Issenhuth-Jeanjean S, Roggentin P, Mikoleit M, Guibourdenche M, de Pinna E, Nair S, Fields PI, Weill FX. Supplement 2008-2010 (no. 48) to the white-Kauffmann-Le minor scheme. Res Microbiol. 2014;165(7):526–530. - PubMed
-
- Keestra-Gounder AM, Tsolis RM, Baumler AJ. Now you see me, now you don't: the interaction of Salmonella with innate immune receptors. Nat Rev Microbiol. 2015;13(4):206–216. - PubMed
-
- Hendriksen RS, Vieira AR, Karlsmose S, Lo Fo Wong DM, Jensen AB, Wegener HC, Aarestrup FM: Global monitoring of Salmonella serovar distribution from the World Health Organization global foodborne infections network country data Bank: results of quality assured laboratories from 2001 to 2007. Foodborne Pathog Dis 2011, 8(8):887–900. - PubMed
MeSH terms
Substances
Grants and funding
- BIO2016-77639-P (AEI/FEDER,UE)/Ministerio de Economía, Industria y Competitividad, Gobierno de España
- PCIN-2016-082/Ministerio de Economía, Industria y Competitividad, Gobierno de España
- 215RT-0493/Programa Iberoamericano de Ciencia y Tecnología para el Desarrollo (CYTED)
- 239659/Consejo Nacional de Ciencia y Tecnología
- FC-2015-2/879/Consejo Nacional de Ciencia y Tecnología
- IN213516/Dirección General de Asuntos del Personal Académico, Universidad Nacional Autónoma de México
- IN211814/Dirección General de Asuntos del Personal Académico, Universidad Nacional Autónoma de México
- IN206318/Dirección General de Asuntos del Personal Académico, Universidad Nacional Autónoma de México

