The AAA+ ATPase/ubiquitin ligase mysterin stabilizes cytoplasmic lipid droplets
- PMID: 30705059
- PMCID: PMC6400562
- DOI: 10.1083/jcb.201712120
The AAA+ ATPase/ubiquitin ligase mysterin stabilizes cytoplasmic lipid droplets
Abstract
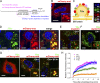
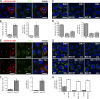
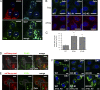
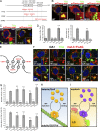
Mysterin, also known as RNF213, is an intracellular protein that forms large toroidal oligomers. Mysterin was originally identified in genetic studies of moyamoya disease (MMD), a rare cerebrovascular disorder of unknown etiology. While mysterin is known to exert ubiquitin ligase and putative mechanical ATPase activities with a RING finger domain and two adjacent AAA+ modules, its biological role is poorly understood. Here, we report that mysterin is targeted to lipid droplets (LDs), ubiquitous organelles specialized for neutral lipid storage, and markedly increases their abundance in cells. This effect was exerted primarily through specific elimination of adipose triglyceride lipase (ATGL) from LDs. The ubiquitin ligase and ATPase activities of mysterin were both important for its proper LD targeting. Notably, MMD-related mutations in the ubiquitin ligase domain of mysterin significantly impaired its fat-stabilizing activity. Our findings identify a unique new regulator of cytoplasmic LDs and suggest a potential link between the pathogenesis of MMD and fat metabolism.
© 2019 Sugihara et al.
Figures
References
-
- Banh R.S., Iorio C., Marcotte R., Xu Y., Cojocari D., Rahman A.A., Pawling J., Zhang W., Sinha A., Rose C.M., et al. . 2016. PTP1B controls non-mitochondrial oxygen consumption by regulating RNF213 to promote tumour survival during hypoxia. Nat. Cell Biol. 18:803–813. 10.1038/ncb3376 - DOI - PMC - PubMed
-
- Bhandari S., Lee J.N., Kim Y.-I., Nam I.-K., Kim S.-J., Kim S.-J., Kwak S., Oh G.-S., Kim H.-J., Yoo H.J., et al. . 2016. The fatty acid chain elongase, Elovl1, is required for kidney and swim bladder development during zebrafish embryogenesis. Organogenesis. 12:78–93. 10.1080/15476278.2016.1172164 - DOI - PMC - PubMed
-
- Cecchi A.C., Guo D., Ren Z., Flynn K., Santos-Cortez R.L.P., Leal S.M., Wang G.T., Regalado E.S., Steinberg G.K., Shendure J., et al. University of Washington Center for Mendelian Genomics . 2014. RNF213 rare variants in an ethnically diverse population with Moyamoya disease. Stroke. 45:3200–3207. 10.1161/STROKEAHA.114.006244 - DOI - PMC - PubMed
-
- Guey S., Kraemer M., Hervé D., Ludwig T., Kossorotoff M., Bergametti F., Schwitalla J.C., Choi S., Broseus L., Callebaut I., et al. ; FREX Consortium . 2017. Rare RNF213 variants in the C-terminal region encompassing the RING-finger domain are associated with moyamoya angiopathy in Caucasians. Eur. J. Hum. Genet. 25:995–1003. 10.1038/ejhg.2017.92 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

